AI 基础:Scipy(科学计算库) 简易入门
0.导语
Scipy是一个用于数学、科学、工程领域的常用软件包,可以处理插值、积分、优化、图像处理、常微分方程数值解的求解、信号处理等问题。它用于有效计算Numpy矩阵,使Numpy和Scipy协同工作,高效解决问题。
Scipy是由针对特定任务的子模块组成:
| 模块名 | 应用领域 |
|---|---|
| scipy.cluster | 向量计算/Kmeans |
| scipy.constants | 物理和数学常量 |
| scipy.fftpack | 傅立叶变换 |
| scipy.integrate | 积分程序 |
| scipy.interpolate | 插值 |
| scipy.io | 数据输入输出 |
| scipy.linalg | 线性代数程序 |
| scipy.ndimage | n维图像包 |
| scipy.odr | 正交距离回归 |
| scipy.optimize | 优化 |
| scipy.signal | 信号处理 |
| scipy.sparse | 稀疏矩阵 |
| scipy.spatial | 空间数据结构和算法 |
| scipy.special | 一些特殊的数学函数 |
| scipy.stats | 统计 |
在此之前,我已经写了以下几篇AI基础的快速入门,本篇文章讲解科学计算库Scipy快速入门:
已发布:
AI 基础:Numpy 简易入门
AI 基础:Pandas 简易入门
AI基础:数据可视化简易入门(matplotlib和seaborn)
备注:本文代码可以在github下载
https://github.com/fengdu78/Data-Science-Notes/tree/master/4.scipy
1.SciPy-数值计算库
import numpy as np
import pylab as pl
import matplotlib as mpl
mpl.rcParams['font.sans-serif'] = ['SimHei']
import scipy
scipy.__version__#查看版本
'1.0.0'
常数和特殊函数
from scipy import constants as C
print (C.c) # 真空中的光速
print (C.h) # 普朗克常数
299792458.0
6.62607004e-34
C.physical_constants["electron mass"]
(9.10938356e-31, 'kg', 1.1e-38)
# 1英里等于多少米, 1英寸等于多少米, 1克等于多少千克, 1磅等于多少千克
print(C.mile)
print(C.inch)
print(C.gram)
print(C.pound)
1609.3439999999998
0.0254
0.001
0.45359236999999997
import scipy.special as S
print (1 + 1e-20)
print (np.log(1+1e-20))
print (S.log1p(1e-20))
1.0
0.0
1e-20
m = np.linspace(0.1, 0.9, 4)
u = np.linspace(-10, 10, 200)
results = S.ellipj(u[:, None], m[None, :])
print([y.shape for y in results])
[(200, 4), (200, 4), (200, 4), (200, 4)]
#%figonly=使用广播计算得到的`ellipj()`返回值
fig, axes = pl.subplots(2, 2, figsize=(12, 4))
labels = ["$sn$", "$cn$", "$dn$", "$\phi$"]
for ax, y, label in zip(axes.ravel(), results, labels):
ax.plot(u, y)
ax.set_ylabel(label)
ax.margins(0, 0.1)
axes[1, 1].legend(["$m={:g}$".format(m_) for m_ in m], loc="best", ncol=2);

2.拟合与优化-optimize
非线性方程组求解
import pylab as pl
import numpy as np
import matplotlib as mpl
mpl.rcParams['font.sans-serif'] = ['SimHei']
from math import sin, cos
from scipy import optimize
def f(x): #❶
x0, x1, x2 = x.tolist() #❷
return [
5*x1+3,
4*x0*x0 - 2*sin(x1*x2),
x1*x2 - 1.5
]
# f计算方程组的误差,[1,1,1]是未知数的初始值
result = optimize.fsolve(f, [1,1,1]) #❸
print (result)
print (f(result))
[-0.70622057 -0.6 -2.5 ]
[0.0, -9.126033262418787e-14, 5.329070518200751e-15]
def j(x): #❶
x0, x1, x2 = x.tolist()
return [[0, 5, 0],
[8 * x0, -2 * x2 * cos(x1 * x2), -2 * x1 * cos(x1 * x2)],
[0, x2, x1]]
result = optimize.fsolve(f, [1, 1, 1], fprime=j) #❷
print(result)
print(f(result))
[-0.70622057 -0.6 -2.5 ]
[0.0, -9.126033262418787e-14, 5.329070518200751e-15]
最小二乘拟合
import numpy as np
from scipy import optimize
X = np.array([ 8.19, 2.72, 6.39, 8.71, 4.7 , 2.66, 3.78])
Y = np.array([ 7.01, 2.78, 6.47, 6.71, 4.1 , 4.23, 4.05])
def residuals(p): #❶
"计算以p为参数的直线和原始数据之间的误差"
k, b = p
return Y - (k*X + b)
# leastsq使得residuals()的输出数组的平方和最小,参数的初始值为[1,0]
r = optimize.leastsq(residuals, [1, 0]) #❷
k, b = r[0]
print ("k =",k, "b =",b)
k = 0.6134953491930442 b = 1.794092543259387
#%figonly=最小化正方形面积之和(左),误差曲面(右)
scale_k = 1.0
scale_b = 10.0
scale_error = 1000.0
def S(k, b):
"计算直线y=k*x+b和原始数据X、Y的误差的平方和"
error = np.zeros(k.shape)
for x, y in zip(X, Y):
error += (y - (k * x + b)) ** 2
return error
ks, bs = np.mgrid[k - scale_k:k + scale_k:40j, b - scale_b:b + scale_b:40j]
error = S(ks, bs) / scale_error
from mpl_toolkits.mplot3d import Axes3D
from matplotlib.patches import Rectangle
fig = pl.figure(figsize=(12, 5))
ax1 = pl.subplot(121)
ax1.plot(X, Y, "o")
X0 = np.linspace(2, 10, 3)
Y0 = k*X0 + b
ax1.plot(X0, Y0)
for x, y in zip(X, Y):
y2 = k*x+b
rect = Rectangle((x,y), abs(y-y2), y2-y, facecolor="red", alpha=0.2)
ax1.add_patch(rect)
ax1.set_aspect("equal")
ax2 = fig.add_subplot(122, projection='3d')
ax2.plot_surface(
ks, bs / scale_b, error, rstride=3, cstride=3, cmap="jet", alpha=0.5)
ax2.scatter([k], [b / scale_b], [S(k, b) / scale_error], c="r", s=20)
ax2.set_xlabel("$k$")
ax2.set_ylabel("$b$")
ax2.set_zlabel("$error$");

#%fig=带噪声的正弦波拟合
def func(x, p): #❶
"""
数据拟合所用的函数: A*sin(2*pi*k*x + theta)
"""
A, k, theta = p
return A * np.sin(2 * np.pi * k * x + theta)
def residuals(p, y, x): #❷
"""
实验数据x, y和拟合函数之间的差,p为拟合需要找到的系数
"""
return y - func(x, p)
x = np.linspace(0, 2 * np.pi, 100)
A, k, theta = 10, 0.34, np.pi / 6 # 真实数据的函数参数
y0 = func(x, [A, k, theta]) # 真实数据
# 加入噪声之后的实验数据
np.random.seed(0)
y1 = y0 + 2 * np.random.randn(len(x)) #❸
p0 = [7, 0.40, 0] # 第一次猜测的函数拟合参数
# 调用leastsq进行数据拟合
# residuals为计算误差的函数
# p0为拟合参数的初始值
# args为需要拟合的实验数据
plsq = optimize.leastsq(residuals, p0, args=(y1, x)) #❹
print(u"真实参数:", [A, k, theta])
print(u"拟合参数", plsq[0]) # 实验数据拟合后的参数
pl.plot(x, y1, "o", label=u"带噪声的实验数据")
pl.plot(x, y0, label=u"真实数据")
pl.plot(x, func(x, plsq[0]), label=u"拟合数据")
pl.legend(loc="best")
真实参数: [10, 0.34, 0.5235987755982988]
拟合参数 [10.25218748 0.3423992 0.50817423]

def func2(x, A, k, theta):
return A*np.sin(2*np.pi*k*x+theta)
popt, _ = optimize.curve_fit(func2, x, y1, p0=p0)
print (popt)
[10.25218748 0.3423992 0.50817425]
popt, _ = optimize.curve_fit(func2, x, y1, p0=[10, 1, 0])
print(u"真实参数:", [A, k, theta])
print(u"拟合参数", popt)
真实参数: [10, 0.34, 0.5235987755982988]
拟合参数 [ 0.71093469 1.02074585 -0.12776742]
计算函数局域最小值
def target_function(x, y):
return (1 - x)**2 + 100 * (y - x**2)**2
class TargetFunction(object):
def __init__(self):
self.f_points = []
self.fprime_points = []
self.fhess_points = []
def f(self, p):
x, y = p.tolist()
z = target_function(x, y)
self.f_points.append((x, y))
return z
def fprime(self, p):
x, y = p.tolist()
self.fprime_points.append((x, y))
dx = -2 + 2 * x - 400 * x * (y - x**2)
dy = 200 * y - 200 * x**2
return np.array([dx, dy])
def fhess(self, p):
x, y = p.tolist()
self.fhess_points.append((x, y))
return np.array([[2 * (600 * x**2 - 200 * y + 1), -400 * x],
[-400 * x, 200]])
def fmin_demo(method):
target = TargetFunction()
init_point = (-1, -1)
res = optimize.minimize(
target.f,
init_point,
method=method,
jac=target.fprime,
hess=target.fhess)
return res, [
np.array(points) for points in (target.f_points, target.fprime_points,
target.fhess_points)
]
methods = ("Nelder-Mead", "Powell", "CG", "BFGS", "Newton-CG", "L-BFGS-B")
for method in methods:
res, (f_points, fprime_points, fhess_points) = fmin_demo(method)
print(
"{:12s}: min={:12g}, f count={:3d}, fprime count={:3d}, fhess count={:3d}"
.format(method, float(res["fun"]), len(f_points), len(fprime_points),
len(fhess_points)))
Nelder-Mead : min= 5.30934e-10, f count=125, fprime count= 0, fhess count= 0
Powell : min= 0, f count= 52, fprime count= 0, fhess count= 0
CG : min= 9.63056e-21, f count= 39, fprime count= 39, fhess count= 0
BFGS : min= 1.84992e-16, f count= 40, fprime count= 40, fhess count= 0
Newton-CG : min= 5.22666e-10, f count= 60, fprime count= 97, fhess count= 38
L-BFGS-B : min= 6.5215e-15, f count= 33, fprime count= 33, fhess count= 0
#%figonly=各种优化算法的搜索路径
def draw_fmin_demo(f_points, fprime_points, ax):
xmin, xmax = -3, 3
ymin, ymax = -3, 3
Y, X = np.ogrid[ymin:ymax:500j,xmin:xmax:500j]
Z = np.log10(target_function(X, Y))
zmin, zmax = np.min(Z), np.max(Z)
ax.imshow(Z, extent=(xmin,xmax,ymin,ymax), origin="bottom", aspect="auto", cmap="gray")
ax.plot(f_points[:,0], f_points[:,1], lw=1)
ax.scatter(f_points[:,0], f_points[:,1], c=range(len(f_points)), s=50, linewidths=0)
if len(fprime_points):
ax.scatter(fprime_points[:, 0], fprime_points[:, 1], marker="x", color="w", alpha=0.5)
ax.set_xlim(xmin, xmax)
ax.set_ylim(ymin, ymax)
fig, axes = pl.subplots(2, 3, figsize=(9, 6))
methods = ("Nelder-Mead", "Powell", "CG", "BFGS", "Newton-CG", "L-BFGS-B")
for ax, method in zip(axes.ravel(), methods):
res, (f_points, fprime_points, fhess_points) = fmin_demo(method)
draw_fmin_demo(f_points, fprime_points, ax)
ax.set_aspect("equal")
ax.set_title(method)

计算全域最小值
def func(x, p):
A, k, theta = p
return A*np.sin(2*np.pi*k*x+theta)
def func_error(p, y, x):
return np.sum((y - func(x, p))**2)
x = np.linspace(0, 2*np.pi, 100)
A, k, theta = 10, 0.34, np.pi/6
y0 = func(x, [A, k, theta])
np.random.seed(0)
y1 = y0 + 2 * np.random.randn(len(x))
result = optimize.basinhopping(func_error, (1, 1, 1),
niter = 10,
minimizer_kwargs={"method":"L-BFGS-B",
"args":(y1, x)})
print (result.x)
[10.25218676 -0.34239909 2.63341581]
#%figonly=用`basinhopping()`拟合正弦波
pl.plot(x, y1, "o", label=u"带噪声的实验数据")
pl.plot(x, y0, label=u"真实数据")
pl.plot(x, func(x, result.x), label=u"拟合数据")
pl.legend(loc="best");

3.线性代数-linalg
解线性方程组
import pylab as pl
import numpy as np
from scipy import linalg
import matplotlib as mpl
mpl.rcParams['font.sans-serif'] = ['SimHei']
import numpy as np
from scipy import linalg
m, n = 500, 50
A = np.random.rand(m, m)
B = np.random.rand(m, n)
X1 = linalg.solve(A, B)
X2 = np.dot(linalg.inv(A), B)
print (np.allclose(X1, X2))
%timeit linalg.solve(A, B)
%timeit np.dot(linalg.inv(A), B)
True
5.38 ms ± 120 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)
8.14 ms ± 994 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)
luf = linalg.lu_factor(A)
X3 = linalg.lu_solve(luf, B)
np.allclose(X1, X3)
True
M, N = 1000, 100
np.random.seed(0)
A = np.random.rand(M, M)
B = np.random.rand(M, N)
Ai = linalg.inv(A)
luf = linalg.lu_factor(A)
%timeit linalg.inv(A)
%timeit np.dot(Ai, B)
%timeit linalg.lu_factor(A)
%timeit linalg.lu_solve(luf, B)
50.6 ms ± 1.94 ms per loop (mean ± std. dev. of 7 runs, 10 loops each)
3.49 ms ± 306 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)
20.1 ms ± 1.42 ms per loop (mean ± std. dev. of 7 runs, 10 loops each)
4.49 ms ± 65 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)
最小二乘解
from numpy.lib.stride_tricks import as_strided
def make_data(m, n, noise_scale): #❶
np.random.seed(42)
x = np.random.standard_normal(m)
h = np.random.standard_normal(n)
y = np.convolve(x, h)
yn = y + np.random.standard_normal(len(y)) * noise_scale * np.max(y)
return x, yn, h
def solve_h(x, y, n): #❷
X = as_strided(
x, shape=(len(x) - n + 1, n), strides=(x.itemsize, x.itemsize)) #❸
Y = y[n - 1:len(x)] #❹
h = linalg.lstsq(X, Y) #❺
return h[0][::-1] #❻
x, yn, h = make_data(1000, 100, 0.4)
H1 = solve_h(x, yn, 120)
H2 = solve_h(x, yn, 80)
print("Average error of H1:", np.mean(np.abs(h[:100] - h)))
print("Average error of H2:", np.mean(np.abs(h[:80] - H2)))
Average error of H1: 0.0
Average error of H2: 0.2958422158342371
#%figonly=实际的系统参数与最小二乘解的比较
fig, (ax1, ax2) = pl.subplots(2, 1, figsize=(6, 4))
ax1.plot(h, linewidth=2, label=u"实际的系统参数")
ax1.plot(H1, linewidth=2, label=u"最小二乘解H1", alpha=0.7)
ax1.legend(loc="best", ncol=2)
ax1.set_xlim(0, len(H1))
ax2.plot(h, linewidth=2, label=u"实际的系统参数")
ax2.plot(H2, linewidth=2, label=u"最小二乘解H2", alpha=0.7)
ax2.legend(loc="best", ncol=2)
ax2.set_xlim(0, len(H1));

特征值和特征向量
A = np.array([[1, -0.3], [-0.1, 0.9]])
evalues, evectors = linalg.eig(A)
print(evalues)
print(evectors)
[1.13027756+0.j 0.76972244+0.j]
[[ 0.91724574 0.79325185]
[-0.3983218 0.60889368]]
#%figonly=线性变换将蓝色箭头变换为红色箭头
points = np.array([[0, 1.0], [1.0, 0], [1, 1]])
def draw_arrows(points, **kw):
props = dict(color="blue", arrowstyle="->")
props.update(kw)
for x, y in points:
pl.annotate("",
xy=(x, y), xycoords='data',
xytext=(0, 0), textcoords='data',
arrowprops=props)
draw_arrows(points)
draw_arrows(np.dot(A, points.T).T, color="red")
draw_arrows(evectors.T, alpha=0.7, linewidth=2)
draw_arrows(np.dot(A, evectors).T, color="red", alpha=0.7, linewidth=2)
ax = pl.gca()
ax.set_aspect("equal")
ax.set_xlim(-0.5, 1.1)
ax.set_ylim(-0.5, 1.1);

np.random.seed(42)
t = np.random.uniform(0, 2*np.pi, 60)
alpha = 0.4
a = 0.5
b = 1.0
x = 1.0 + a*np.cos(t)*np.cos(alpha) - b*np.sin(t)*np.sin(alpha)
y = 1.0 + a*np.cos(t)*np.sin(alpha) - b*np.sin(t)*np.cos(alpha)
x += np.random.normal(0, 0.05, size=len(x))
y += np.random.normal(0, 0.05, size=len(y))
D = np.c_[x**2, x*y, y**2, x, y, np.ones_like(x)]
A = np.dot(D.T, D)
C = np.zeros((6, 6))
C[[0, 1, 2], [2, 1, 0]] = 2, -1, 2
evalues, evectors = linalg.eig(A, C) #❶
evectors = np.real(evectors)
err = np.mean(np.dot(D, evectors)**2, 0) #❷
p = evectors[:, np.argmin(err) ] #❸
print (p)
[-0.55214278 0.5580915 -0.23809922 0.54584559 -0.08350449 -0.14852803]
#%figonly=用广义特征向量计算的拟合椭圆
def ellipse(p, x, y):
a, b, c, d, e, f = p
return a*x**2 + b*x*y + c*y**2 + d*x + e*y + f
X, Y = np.mgrid[0:2:100j, 0:2:100j]
Z = ellipse(p, X, Y)
pl.plot(x, y, "ro", alpha=0.5)
pl.contour(X, Y, Z, levels=[0]);

奇异值分解-SVD
r, g, b = np.rollaxis(pl.imread("vinci_target.png"), 2).astype(float)
img = 0.2989 * r + 0.5870 * g + 0.1140 * b
img.shape
(505, 375)
U, s, Vh = linalg.svd(img)
print(U.shape)
print(s.shape)
print(Vh.shape)
(505, 505)
(375,)
(375, 375)
#%fig=按从大到小排列的奇异值
pl.semilogy(s, lw=3);
 output_20_1
output_20_1
def composite(U, s, Vh, n):
return np.dot(U[:, :n], s[:n, np.newaxis] * Vh[:n, :])
print (np.allclose(img, composite(U, s, Vh, len(s))))
True
#%fig=原始图像、使用10、20、50个向量合成的图像(从左到右)
img10 = composite(U, s, Vh, 10)
img20 = composite(U, s, Vh, 20)
img50 = composite(U, s, Vh, 50)
%array_image img; img10; img20; img50
UsageError: Line magic function `%array_image` not found.
pl.imshow(img)

pl.imshow(img10)

pl.imshow(img20)

pl.imshow(img50)

4.统计-stats
import numpy as np
import pylab as pl
from scipy import stats
import matplotlib.pyplot as plt
import matplotlib as mpl
mpl.rcParams['font.sans-serif'] = ['SimHei']
连续概率分布
from scipy import stats
[k for k, v in stats.__dict__.items() if isinstance(v, stats.rv_continuous)]
['ksone',
'kstwobign',
'norm',
'alpha',
'anglit',
'arcsine',
'beta',
'betaprime',
'bradford',
'burr',
'burr12',
'fisk',
'cauchy',
'chi',
'chi2',
'cosine',
'dgamma',
'dweibull',
'expon',
'exponnorm',
'exponweib',
'exponpow',
'fatiguelife',
'foldcauchy',
'f',
'foldnorm',
'weibull_min',
'weibull_max',
'frechet_r',
'frechet_l',
'genlogistic',
'genpareto',
'genexpon',
'genextreme',
'gamma',
'erlang',
'gengamma',
'genhalflogistic',
'gompertz',
'gumbel_r',
'gumbel_l',
'halfcauchy',
'halflogistic',
'halfnorm',
'hypsecant',
'gausshyper',
'invgamma',
'invgauss',
'invweibull',
'johnsonsb',
'johnsonsu',
'laplace',
'levy',
'levy_l',
'levy_stable',
'logistic',
'loggamma',
'loglaplace',
'lognorm',
'gilbrat',
'maxwell',
'mielke',
'kappa4',
'kappa3',
'nakagami',
'ncx2',
'ncf',
't',
'nct',
'pareto',
'lomax',
'pearson3',
'powerlaw',
'powerlognorm',
'powernorm',
'rdist',
'rayleigh',
'reciprocal',
'rice',
'recipinvgauss',
'semicircular',
'skewnorm',
'trapz',
'triang',
'truncexpon',
'truncnorm',
'tukeylambda',
'uniform',
'vonmises',
'vonmises_line',
'wald',
'wrapcauchy',
'gennorm',
'halfgennorm',
'crystalball',
'argus']
stats.norm.stats()
(array(0.), array(1.))
X = stats.norm(loc=1.0, scale=2.0)
X.stats()
(array(1.), array(4.))
x = X.rvs(size=10000) # 对随机变量取10000个值
np.mean(x), np.var(x) # 期望值和方差
(1.0048352738823323, 3.9372117720073554)
stats.norm.fit(x) # 得到随机序列期望值和标准差
(1.0048352738823323, 1.984240855341749)
pdf, t = np.histogram(x, bins=100, normed=True) #❶
t = (t[:-1] + t[1:]) * 0.5 #❷
cdf = np.cumsum(pdf) * (t[1] - t[0]) #❸
p_error = pdf - X.pdf(t)
c_error = cdf - X.cdf(t)
print ("max pdf error: {}, max cdf error: {}".format(
np.abs(p_error).max(),
np.abs(c_error).max()))
max pdf error: 0.018998755595167102, max cdf error: 0.018503342378306975
#%figonly=正态分布的概率密度函数(左)和累积分布函数(右)
fig, (ax1, ax2) = pl.subplots(1, 2, figsize=(7, 2))
ax1.plot(t, pdf, label=u"统计值")
ax1.plot(t, X.pdf(t), label=u"理论值", alpha=0.6)
ax1.legend(loc="best")
ax2.plot(t, cdf)
ax2.plot(t, X.cdf(t), alpha=0.6);

print(stats.gamma.stats(1.0))
print(stats.gamma.stats(2.0))
(array(1.), array(1.))
(array(2.), array(2.))
stats.gamma.stats(2.0, scale=2)
(array(4.), array(8.))
x = stats.gamma.rvs(2, scale=2, size=4)
x
array([4.40563983, 6.17699951, 3.65503843, 3.28052152])
stats.gamma.pdf(x, 2, scale=2)
array([0.12169605, 0.07037188, 0.14694352, 0.15904745])
X = stats.gamma(2, scale=2)
X.pdf(x)
array([0.12169605, 0.07037188, 0.14694352, 0.15904745])
离散概率分布
x = range(1, 7)
p = (0.4, 0.2, 0.1, 0.1, 0.1, 0.1)
dice = stats.rv_discrete(values=(x, p))
dice.rvs(size=20)
array([2, 5, 2, 6, 1, 6, 6, 5, 3, 1, 5, 2, 1, 1, 1, 1, 1, 2, 1, 6])
np.random.seed(42)
samples = dice.rvs(size=(20000, 50))
samples_mean = np.mean(samples, axis=1)
核密度估计
#%fig=核密度估计能更准确地表示随机变量的概率密度函数
_, bins, step = pl.hist(
samples_mean, bins=100, normed=True, histtype="step", label=u"直方图统计")
kde = stats.kde.gaussian_kde(samples_mean)
x = np.linspace(bins[0], bins[-1], 100)
pl.plot(x, kde(x), label=u"核密度估计")
mean, std = stats.norm.fit(samples_mean)
pl.plot(x, stats.norm(mean, std).pdf(x), alpha=0.8, label=u"正态分布拟合")
pl.legend()

#%fig=`bw_method`参数越大核密度估计曲线越平滑
for bw in [0.2, 0.3, 0.6, 1.0]:
kde = stats.gaussian_kde([-1, 0, 1], bw_method=bw)
x = np.linspace(-5, 5, 1000)
y = kde(x)
pl.plot(x, y, lw=2, label="bw={}".format(bw), alpha=0.6)
pl.legend(loc="best");

二项、泊松、伽玛分布
stats.binom.pmf(range(6), 5, 1/6.0)
array([4.01877572e-01, 4.01877572e-01, 1.60751029e-01, 3.21502058e-02,
3.21502058e-03, 1.28600823e-04])
#%fig=当n足够大时二项分布和泊松分布近似相等
lambda_ = 10.0
x = np.arange(20)
n1, n2 = 100, 1000
y_binom_n1 = stats.binom.pmf(x, n1, lambda_ / n1)
y_binom_n2 = stats.binom.pmf(x, n2, lambda_ / n2)
y_poisson = stats.poisson.pmf(x, lambda_)
print(np.max(np.abs(y_binom_n1 - y_poisson)))
print(np.max(np.abs(y_binom_n2 - y_poisson)))
#%hide
fig, (ax1, ax2) = pl.subplots(1, 2, figsize=(7.5, 2.5))
ax1.plot(x, y_binom_n1, label=u"binom", lw=2)
ax1.plot(x, y_poisson, label=u"poisson", lw=2, color="red")
ax2.plot(x, y_binom_n2, label=u"binom", lw=2)
ax2.plot(x, y_poisson, label=u"poisson", lw=2, color="red")
for n, ax in zip((n1, n2), (ax1, ax2)):
ax.set_xlabel(u"次数")
ax.set_ylabel(u"概率")
ax.set_title("n={}".format(n))
ax.legend()
fig.subplots_adjust(0.1, 0.15, 0.95, 0.90, 0.2, 0.1)
0.00675531110335309
0.0006301754049777564
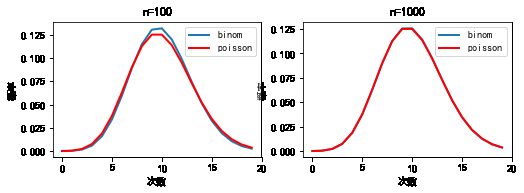
#%fig=模拟泊松分布
np.random.seed(42)
def sim_poisson(lambda_, time):
t = np.random.uniform(0, time, size=lambda_ * time) #❶
count, time_edges = np.histogram(t, bins=time, range=(0, time)) #❷
dist, count_edges = np.histogram(
count, bins=20, range=(0, 20), density=True) #❸
x = count_edges[:-1]
poisson = stats.poisson.pmf(x, lambda_)
return x, poisson, dist
lambda_ = 10
times = 1000, 50000
x1, poisson1, dist1 = sim_poisson(lambda_, times[0])
x2, poisson2, dist2 = sim_poisson(lambda_, times[1])
max_error1 = np.max(np.abs(dist1 - poisson1))
max_error2 = np.max(np.abs(dist2 - poisson2))
print("time={}, max_error={}".format(times[0], max_error1))
print("time={}, max_error={}".format(times[1], max_error2))
#%hide
fig, (ax1, ax2) = pl.subplots(1, 2, figsize=(7.5, 2.5))
ax1.plot(x1, dist1, "-o", lw=2, label=u"统计结果")
ax1.plot(x1, poisson1, "->", lw=2, label=u"泊松分布", color="red", alpha=0.6)
ax2.plot(x2, dist2, "-o", lw=2, label=u"统计结果")
ax2.plot(x2, poisson2, "->", lw=2, label=u"泊松分布", color="red", alpha=0.6)
for ax, time in zip((ax1, ax2), times):
ax.set_xlabel(u"次数")
ax.set_ylabel(u"概率")
ax.set_title(u"time = {}".format(time))
ax.legend(loc="lower center")
fig.subplots_adjust(0.1, 0.15, 0.95, 0.90, 0.2, 0.1)
time=1000, max_error=0.01964230201602718
time=50000, max_error=0.001798012894964722

#%fig=模拟伽玛分布
def sim_gamma(lambda_, time, k):
t = np.random.uniform(0, time, size=lambda_ * time) #❶
t.sort() #❷
interval = t[k:] - t[:-k] #❸
dist, interval_edges = np.histogram(interval, bins=100, density=True) #❹
x = (interval_edges[1:] + interval_edges[:-1])/2 #❺
gamma = stats.gamma.pdf(x, k, scale=1.0/lambda_) #❺
return x, gamma, dist
lambda_ = 10
time = 1000
ks = 1, 2
x1, gamma1, dist1 = sim_gamma(lambda_, time, ks[0])
x2, gamma2, dist2 = sim_gamma(lambda_, time, ks[1])
#%hide
fig, (ax1, ax2) = pl.subplots(1, 2, figsize=(7.5, 2.5))
ax1.plot(x1, dist1, lw=2, label=u"统计结果")
ax1.plot(x1, gamma1, lw=2, label=u"伽玛分布", color="red", alpha=0.6)
ax2.plot(x2, dist2, lw=2, label=u"统计结果")
ax2.plot(x2, gamma2, lw=2, label=u"伽玛分布", color="red", alpha=0.6)
for ax, k in zip((ax1, ax2), ks):
ax.set_xlabel(u"时间间隔")
ax.set_ylabel(u"概率密度")
ax.set_title(u"k = {}".format(k))
ax.legend(loc="upper right")
fig.subplots_adjust(0.1, 0.15, 0.95, 0.90, 0.2, 0.1);
T = 100000
A_count = int(T / 5)
B_count = int(T / 10)
A_time = np.random.uniform(0, T, A_count) #❶
B_time = np.random.uniform(0, T, B_count)
bus_time = np.concatenate((A_time, B_time)) #❷
bus_time.sort()
N = 200000
passenger_time = np.random.uniform(bus_time[0], bus_time[-1], N) #❸
idx = np.searchsorted(bus_time, passenger_time) #❹
np.mean(bus_time[idx] - passenger_time) * 60 #❺
202.3388747879705
np.mean(np.diff(bus_time)) * 60
199.99833251643057
#%figonly=观察者偏差
import matplotlib.gridspec as gridspec
pl.figure(figsize=(7.5, 3))
G = gridspec.GridSpec(10, 1)
ax1 = pl.subplot(G[:2, 0])
ax2 = pl.subplot(G[3:, 0])
ax1.vlines(bus_time[:10], 0, 1, lw=2, color="blue", label=u"公交车")
ptime = np.random.uniform(bus_time[0], bus_time[9], 100)
ax1.vlines(ptime, 0, 1, lw=1, color="red", alpha=0.2, label=u"乘客")
ax1.legend()
count, bins = np.histogram(passenger_time, bins=bus_time)
ax2.plot(np.diff(bins), count, ".", alpha=0.3, rasterized=True)
ax2.set_xlabel(u"公交车的时间间隔")
ax2.set_ylabel(u"等待人数");

from scipy import integrate
t = 10.0 / 3 # 两辆公交车之间的平均时间间隔
bus_interval = stats.gamma(1, scale=t)
n, _ = integrate.quad(lambda x: 0.5 * x * x * bus_interval.pdf(x), 0, 1000)
d, _ = integrate.quad(lambda x: x * bus_interval.pdf(x), 0, 1000)
n / d * 60
200.0
学生 t-分布与 t 检验
#%fig=模拟学生t-分布
mu = 0.0
n = 10
samples = stats.norm(mu).rvs(size=(100000, n)) #❶
t_samples = (np.mean(samples, axis=1) - mu) / np.std(
samples, ddof=1, axis=1) * n**0.5 #❷
sample_dist, x = np.histogram(t_samples, bins=100, density=True) #❸
x = 0.5 * (x[:-1] + x[1:])
t_dist = stats.t(n - 1).pdf(x)
print("max error:", np.max(np.abs(sample_dist - t_dist)))
#%hide
pl.plot(x, sample_dist, lw=2, label=u"样本分布")
pl.plot(x, t_dist, lw=2, alpha=0.6, label=u"t分布")
pl.xlim(-5, 5)
pl.legend(loc="best")
max error: 0.006832108369761447

#%figonly=当`df`增大,学生t-分布趋向于正态分布
fig, (ax1, ax2) = pl.subplots(1, 2, figsize=(7.5, 2.5))
ax1.plot(x, stats.t(6-1).pdf(x), label=u"df=5", lw=2)
ax1.plot(x, stats.t(40-1).pdf(x), label=u"df=39", lw=2, alpha=0.6)
ax1.plot(x, stats.norm.pdf(x), "k--", label=u"norm")
ax1.legend()
ax2.plot(x, stats.t(6-1).sf(x), label=u"df=5", lw=2)
ax2.plot(x, stats.t(40-1).sf(x), label=u"df=39", lw=2, alpha=0.6)
ax2.plot(x, stats.norm.sf(x), "k--", label=u"norm")
ax2.legend();

n = 30
np.random.seed(42)
s = stats.norm.rvs(loc=1, scale=0.8, size=n)
t = (np.mean(s) - 0.5) / (np.std(s, ddof=1) / np.sqrt(n))
print (t, stats.ttest_1samp(s, 0.5))
2.658584340882224 Ttest_1sampResult(statistic=2.658584340882224, pvalue=0.01263770225709123)
print ((np.mean(s) - 1) / (np.std(s, ddof=1) / np.sqrt(n)))
print (stats.ttest_1samp(s, 1), stats.ttest_1samp(s, 0.9))
-1.1450173670383303
Ttest_1sampResult(statistic=-1.1450173670383303, pvalue=0.26156414618801477) Ttest_1sampResult(statistic=-0.3842970254542196, pvalue=0.7035619103425202)
#%fig=红色部分为`ttest_1samp()`计算的p值
x = np.linspace(-5, 5, 500)
y = stats.t(n-1).pdf(x)
plt.plot(x, y, lw=2)
t, p = stats.ttest_1samp(s, 0.5)
mask = x > np.abs(t)
plt.fill_between(x[mask], y[mask], color="red", alpha=0.5)
mask = x < -np.abs(t)
plt.fill_between(x[mask], y[mask], color="red", alpha=0.5)
plt.axhline(color="k", lw=0.5)
plt.xlim(-5, 5);

from scipy import integrate
x = np.linspace(-10, 10, 100000)
y = stats.t(n-1).pdf(x)
mask = x >= np.abs(t)
integrate.trapz(y[mask], x[mask])*2
0.012633433707685974
m = 200000
mean = 0.5
r = stats.norm.rvs(loc=mean, scale=0.8, size=(m, n))
ts = (np.mean(s) - mean) / (np.std(s, ddof=1) / np.sqrt(n))
tr = (np.mean(r, axis=1) - mean) / (np.std(r, ddof=1, axis=1) / np.sqrt(n))
np.mean(np.abs(tr) > np.abs(ts))
0.012695
np.random.seed(42)
s1 = stats.norm.rvs(loc=1, scale=1.0, size=20)
s2 = stats.norm.rvs(loc=1.5, scale=0.5, size=20)
s3 = stats.norm.rvs(loc=1.5, scale=0.5, size=25)
print (stats.ttest_ind(s1, s2, equal_var=False)) #❶
print (stats.ttest_ind(s2, s3, equal_var=True)) #❷
Ttest_indResult(statistic=-2.2391470627176755, pvalue=0.033250866086743665)
Ttest_indResult(statistic=-0.5946698521856172, pvalue=0.5551805875810539)
卡方分布和卡方检验
#%fig=使用随机数验证卡方分布
a = np.random.normal(size=(300000, 4))
cs = np.sum(a**2, axis=1)
sample_dist, bins = np.histogram(cs, bins=100, range=(0, 20), density=True)
x = 0.5 * (bins[:-1] + bins[1:])
chi2_dist = stats.chi2.pdf(x, 4)
print("max error:", np.max(np.abs(sample_dist - chi2_dist)))
#%hide
pl.plot(x, sample_dist, lw=2, label=u"样本分布")
pl.plot(x, chi2_dist, lw=2, alpha=0.6, label=u"$\chi ^{2}$分布")
pl.legend(loc="best")
max error: 0.0030732520533635066

#%fig=模拟卡方分布
repeat_count = 60000
n, k = 100, 5
np.random.seed(42)
ball_ids = np.random.randint(0, k, size=(repeat_count, n)) #❶
counts = np.apply_along_axis(np.bincount, 1, ball_ids, minlength=k) #❷
cs2 = np.sum((counts - n/k)**2.0/(n/k), axis=1) #❸
k = stats.kde.gaussian_kde(cs2) #❹
x = np.linspace(0, 10, 200)
pl.plot(x, stats.chi2.pdf(x, 4), lw=2, label=u"$\chi ^{2}$分布")
pl.plot(x, k(x), lw=2, color="red", alpha=0.6, label=u"样本分布")
pl.legend(loc="best")
pl.xlim(0, 10);

def choose_balls(probabilities, size):
r = stats.rv_discrete(values=(range(len(probabilities)), probabilities))
s = r.rvs(size=size)
counts = np.bincount(s)
return counts
np.random.seed(42)
counts1 = choose_balls([0.18, 0.24, 0.25, 0.16, 0.17], 400)
counts2 = choose_balls([0.2]*5, 400)
print(counts1)
print(counts2)
[80 93 97 64 66]
[89 76 79 71 85]
chi1, p1 = stats.chisquare(counts1)
chi2, p2 = stats.chisquare(counts2)
print ("chi1 =", chi1, "p1 =", p1)
print ("chi2 =", chi2, "p2 =", p2)
chi1 = 11.375 p1 = 0.022657601239769634
chi2 = 2.55 p2 = 0.6357054527037017
#%figonly=卡方检验计算的概率为阴影部分的面积
x = np.linspace(0, 30, 200)
CHI2 = stats.chi2(4)
pl.plot(x, CHI2.pdf(x), "k", lw=2)
pl.vlines(chi1, 0, CHI2.pdf(chi1))
pl.vlines(chi2, 0, CHI2.pdf(chi2))
pl.fill_between(x[x>chi1], 0, CHI2.pdf(x[x>chi1]), color="red", alpha=1.0)
pl.fill_between(x[x>chi2], 0, CHI2.pdf(x[x>chi2]), color="green", alpha=0.5)
pl.text(chi1, 0.015, r"$\chi^2_1$", fontsize=14)
pl.text(chi2, 0.015, r"$\chi^2_2$", fontsize=14)
pl.ylim(0, 0.2)
pl.xlim(0, 20);
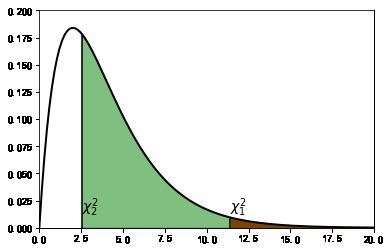
table = [[43, 9], [44, 4]]
chi2, p, dof, expected = stats.chi2_contingency(table)
print(chi2)
print(p)
1.0724852071005921
0.300384770390566
stats.fisher_exact(table)
(0.43434343434343436, 0.23915695682224306)
5.数值积分-integrate
import pylab as pl
import numpy as np
from scipy import integrate
from scipy.integrate import odeint
import matplotlib as mpl
mpl.rcParams['font.sans-serif'] = ['SimHei']
球的体积
def half_circle(x):
return (1-x**2)**0.5
N = 10000
x = np.linspace(-1, 1, N)
dx = x[1] - x[0]
y = half_circle(x)
2 * dx * np.sum(y) # 面积的两倍
3.1415893269307373
np.trapz(y, x) * 2 # 面积的两倍
3.1415893269315975
from scipy import integrate
pi_half, err = integrate.quad(half_circle, -1, 1)
pi_half * 2
3.141592653589797
def half_sphere(x, y):
return (1-x**2-y**2)**0.5
volume, error = integrate.dblquad(half_sphere, -1, 1,
lambda x:-half_circle(x),
lambda x:half_circle(x))
print (volume, error, np.pi*4/3/2)
2.094395102393199 1.0002356720661965e-09 2.0943951023931953
解常微分方程组
#%fig=洛伦茨吸引子:微小的初值差别也会显著地影响运动轨迹
from scipy.integrate import odeint
import numpy as np
def lorenz(w, t, p, r, b): #❶
# 给出位置矢量w,和三个参数p, r, b计算出
# dx/dt, dy/dt, dz/dt的值
x, y, z = w.tolist()
# 直接与lorenz的计算公式对应
return p*(y-x), x*(r-z)-y, x*y-b*z
t = np.arange(0, 30, 0.02) # 创建时间点
# 调用ode对lorenz进行求解, 用两个不同的初始值
track1 = odeint(lorenz, (0.0, 1.00, 0.0), t, args=(10.0, 28.0, 3.0)) #❷
track2 = odeint(lorenz, (0.0, 1.01, 0.0), t, args=(10.0, 28.0, 3.0)) #❸
#%hide
from mpl_toolkits.mplot3d import Axes3D
fig = pl.figure()
ax = Axes3D(fig)
ax.plot(track1[:,0], track1[:,1], track1[:,2], lw=1)
ax.plot(track2[:,0], track2[:,1], track2[:,2], lw=1);
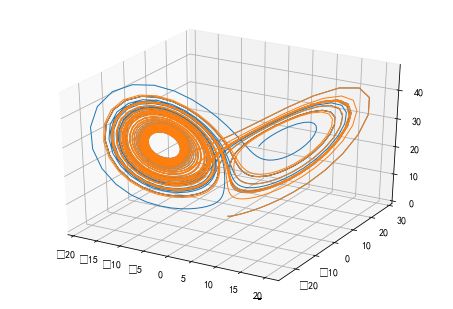
ode 类
def mass_spring_damper(xu, t, m, k, b, F):
x, u = xu.tolist()
dx = u
du = (F - k*x - b*u)/m
return dx, du
#%fig=滑块的速度和位移曲线
m, b, k, F = 1.0, 10.0, 20.0, 1.0
init_status = 0.0, 0.0
args = m, k, b, F
t = np.arange(0, 2, 0.01)
result = odeint(mass_spring_damper, init_status, t, args)
#%hide
fig, (ax1, ax2) = pl.subplots(2, 1)
ax1.plot(t, result[:, 0], label=u"位移")
ax1.legend()
ax2.plot(t, result[:, 1], label=u"速度")
ax2.legend();

from scipy.integrate import ode
class MassSpringDamper(object): #❶
def __init__(self, m, k, b, F):
self.m, self.k, self.b, self.F = m, k, b, F
def f(self, t, xu):
x, u = xu.tolist()
dx = u
du = (self.F - self.k*x - self.b*u)/self.m
return [dx, du]
system = MassSpringDamper(m=m, k=k, b=b, F=F)
init_status = 0.0, 0.0
dt = 0.01
r = ode(system.f) #❷
r.set_integrator('vode', method='bdf')
r.set_initial_value(init_status, 0)
t = []
result2 = [init_status]
while r.successful() and r.t + dt < 2: #❸
r.integrate(r.t + dt)
t.append(r.t)
result2.append(r.y)
result2 = np.array(result2)
np.allclose(result, result2)
True
class PID(object):
def __init__(self, kp, ki, kd, dt):
self.kp, self.ki, self.kd, self.dt = kp, ki, kd, dt
self.last_error = None
self.status = 0.0
def update(self, error):
p = self.kp * error
i = self.ki * self.status
if self.last_error is None:
d = 0.0
else:
d = self.kd * (error - self.last_error) / self.dt
self.status += error * self.dt
self.last_error = error
return p + i + d
#%fig=使用PID控制器让滑块停在位移为1.0处
def pid_control_system(kp, ki, kd, dt, target=1.0):
system = MassSpringDamper(m=m, k=k, b=b, F=0.0)
pid = PID(kp, ki, kd, dt)
init_status = 0.0, 0.0
r = ode(system.f)
r.set_integrator('vode', method='bdf')
r.set_initial_value(init_status, 0)
t = [0]
result = [init_status]
F_arr = [0]
while r.successful() and r.t + dt < 2.0:
r.integrate(r.t + dt)
err = target - r.y[0] #❶
F = pid.update(err) #❷
system.F = F #❸
t.append(r.t)
result.append(r.y)
F_arr.append(F)
result = np.array(result)
t = np.array(t)
F_arr = np.array(F_arr)
return t, F_arr, result
t, F_arr, result = pid_control_system(50.0, 100.0, 10.0, 0.001)
print(u"控制力的终值:", F_arr[-1])
#%hide
fig, (ax1, ax2, ax3) = pl.subplots(3, 1, figsize=(6, 6))
ax1.plot(t, result[:, 0], label=u"位移")
ax1.legend(loc="best")
ax2.plot(t, result[:, 1], label=u"速度")
ax2.legend(loc="best")
ax3.plot(t, F_arr, label=u"控制力")
ax3.legend(loc="best")
控制力的终值: 19.943404699515057

%%time
from scipy import optimize
def eval_func(k):
kp, ki, kd = k
t, F_arr, result = pid_control_system(kp, ki, kd, 0.01)
return np.sum(np.abs(result[:, 0] - 1.0))
kwargs = {
"method": "L-BFGS-B",
"bounds": [(10, 200), (10, 100), (1, 100)],
"options": {
"approx_grad": True
}
}
opt_k = optimize.basinhopping(
eval_func, (10, 10, 10), niter=10, minimizer_kwargs=kwargs)
print(opt_k.x)
[56.67106149 99.97434757 1.33963577]
Wall time: 55.1 s
#%fig=优化PID的参数降低控制响应时间
kp, ki, kd = opt_k.x
t, F_arr, result = pid_control_system(kp, ki, kd, 0.01)
idx = np.argmin(np.abs(t - 0.5))
x, u = result[idx]
print ("t={}, x={:g}, u={:g}".format(t[idx], x, u))
#%hide
fig, (ax1, ax2, ax3) = pl.subplots(3, 1, figsize=(6, 6))
ax1.plot(t, result[:, 0], label=u"位移")
ax1.legend(loc="best")
ax2.plot(t, result[:, 1], label=u"速度")
ax2.legend(loc="best")
ax3.plot(t, F_arr, label=u"控制力")
ax3.legend(loc="best");
t=0.5000000000000002, x=1.07098, u=0.315352
6.信号处理-signal
import pylab as pl
import numpy as np
from scipy import signal
import matplotlib as mpl
mpl.rcParams['font.sans-serif'] = ['SimHei']
中值滤波
#%fig=使用中值滤波剔除瞬间噪声
t = np.arange(0, 20, 0.1)
x = np.sin(t)
x[np.random.randint(0, len(t), 20)] += np.random.standard_normal(20)*0.6 #❶
x2 = signal.medfilt(x, 5) #❷
x3 = signal.order_filter(x, np.ones(5), 2)
print (np.all(x2 == x3))
pl.plot(t, x, label=u"带噪声的信号")
pl.plot(t, x2 + 0.5, alpha=0.6, label=u"中值滤波之后的信号")
pl.legend(loc="best");
True
 output_4_1
output_4_1
滤波器设计
sampling_rate = 8000.0
# 设计一个带通滤波器:
# 通带为0.2*4000 - 0.5*4000
# 阻带为<0.1*4000, >0.6*4000
# 通带增益的最大衰减值为2dB
# 阻带的最小衰减值为40dB
b, a = signal.iirdesign([0.2, 0.5], [0.1, 0.6], 2, 40) #❶
# 使用freq计算滤波器的频率响应
w, h = signal.freqz(b, a) #❷
# 计算增益
power = 20*np.log10(np.clip(np.abs(h), 1e-8, 1e100)) #❸
freq = w / np.pi * sampling_rate / 2
#%fig=用频率扫描波测量的频率响应
# 产生2秒钟的取样频率为sampling_rate Hz的频率扫描信号
# 开始频率为0, 结束频率为sampling_rate/2
t = np.arange(0, 2, 1/sampling_rate) #❶
sweep = signal.chirp(t, f0=0, t1=2, f1=sampling_rate/2) #❷
# 对频率扫描信号进行滤波
out = signal.lfilter(b, a, sweep) #❸
# 将波形转换为能量
out = 20*np.log10(np.abs(out)) #❹
# 找到所有局部最大值的下标
index = signal.argrelmax(out, order=3) #❺
# 绘制滤波之后的波形的增益
pl.figure(figsize=(8, 2.5))
pl.plot(freq, power, label=u"带通IIR滤波器的频率响应")
pl.plot(t[index]/2.0*4000, out[index], label=u"频率扫描波测量的频谱", alpha=0.6) #❻
pl.legend(loc="best")
#%hide
pl.title(u"频率扫描波测量的滤波器频谱")
pl.ylim(-100,20)
pl.ylabel(u"增益(dB)")
pl.xlabel(u"频率(Hz)");

连续时间线性系统
#%fig=系统的阶跃响应和正弦波响应
m, b, k = 1.0, 10, 20
numerator = [1]
denominator = [m, b, k]
plant = signal.lti(numerator, denominator) #❶
t = np.arange(0, 2, 0.01)
_, x_step = plant.step(T=t) #❷
_, x_sin, _ = signal.lsim(plant, U=np.sin(np.pi * t), T=t) #❸
#%hide
pl.plot(t, x_step, label=u"阶跃响应")
pl.plot(t, x_sin, label=u"正弦波响应")
pl.legend(loc="best")
pl.xlabel(u"时间(秒)")
pl.ylabel(u"位移(米)")
Text(0,0.5,'位移(米)')

7.插值-interpolate
import numpy as np
import pylab as pl
from scipy import interpolate
import matplotlib as mpl
mpl.rcParams['font.sans-serif'] = ['SimHei']
一维插值
WARNING:高次
interp1d()插值的运算量很大,因此对于点数较多的数据,建议使用后面介绍的UnivariateSpline()。
#%fig=`interp1d`的各阶插值
from scipy import interpolate
x = np.linspace(0, 10, 11)
y = np.sin(x)
xnew = np.linspace(0, 10, 101)
pl.plot(x, y, 'ro')
for kind in ['nearest', 'zero', 'slinear', 'quadratic']:
f = interpolate.interp1d(x, y, kind=kind) #❶
ynew = f(xnew) #❷
pl.plot(xnew, ynew, label=str(kind))
pl.legend(loc='lower right')
 output_5_1
output_5_1
外推和 Spline 拟合
#%fig=使用UnivariateSpline进行插值:外推(上),数据拟合(下)
x1 = np.linspace(0, 10, 20)
y1 = np.sin(x1)
sx1 = np.linspace(0, 12, 100)
sy1 = interpolate.UnivariateSpline(x1, y1, s=0)(sx1) #❶
x2 = np.linspace(0, 20, 200)
y2 = np.sin(x2) + np.random.standard_normal(len(x2)) * 0.2
sx2 = np.linspace(0, 20, 2000)
spline2 = interpolate.UnivariateSpline(x2, y2, s=8) #❷
sy2 = spline2(sx2)
pl.figure(figsize=(8, 5))
pl.subplot(211)
pl.plot(x1, y1, ".", label=u"数据点")
pl.plot(sx1, sy1, label=u"spline曲线")
pl.legend()
pl.subplot(212)
pl.plot(x2, y2, ".", label=u"数据点")
pl.plot(sx2, sy2, linewidth=2, label=u"spline曲线")
pl.plot(x2, np.sin(x2), label=u"无噪声曲线")
pl.legend()
 output_7_1
output_7_1
print(np.array_str(spline2.roots(), precision=3))
[ 0.053 3.151 6.36 9.386 12.603 15.619 18.929]
#%fig=计算Spline与水平线的交点
def roots_at(self, v): #❶
coeff = self.get_coeffs()
coeff -= v
try:
root = self.roots()
return root
finally:
coeff += v
interpolate.UnivariateSpline.roots_at = roots_at #❷
pl.plot(sx2, sy2, linewidth=2, label=u"spline曲线")
ax = pl.gca()
for level in [0.5, 0.75, -0.5, -0.75]:
ax.axhline(level, ls=":", color="k")
xr = spline2.roots_at(level) #❸
pl.plot(xr, spline2(xr), "ro")
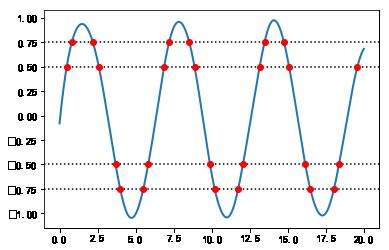
参数插值
#%fig=使用参数插值连接二维平面上的点
x = [
4.913, 4.913, 4.918, 4.938, 4.955, 4.949, 4.911, 4.848, 4.864, 4.893,
4.935, 4.981, 5.01, 5.021
]
y = [
5.2785, 5.2875, 5.291, 5.289, 5.28, 5.26, 5.245, 5.245, 5.2615, 5.278,
5.2775, 5.261, 5.245, 5.241
]
pl.plot(x, y, "o")
for s in (0, 1e-4):
tck, t = interpolate.splprep([x, y], s=s) #❶
xi, yi = interpolate.splev(np.linspace(t[0], t[-1], 200), tck) #❷
pl.plot(xi, yi, lw=2, label=u"s=%g" % s)
pl.legend()

单调插值
import numpy as np
import matplotlib.pyplot as plt
from scipy import interpolate
x = np.arange(0, 2 * np.pi + np.pi / 4, 2 * np.pi / 8)
y = np.sin(x)
tck = interpolate.splrep(x, y, s=0)
xnew = np.arange(0, 2 * np.pi, np.pi / 50)
ynew = interpolate.splev(xnew, tck, der=0)
plt.figure()
plt.plot(x, y, 'x', xnew, ynew, xnew, np.sin(xnew), x, y, 'b')
plt.legend(['Linear', 'Cubic Spline', 'True'])
plt.axis([-0.05, 6.33, -1.05, 1.05])
plt.title('三次样条插值')
plt.show()

多维插值
#%fig=使用interp2d类进行二维插值
def func(x, y): #❶
return (x + y) * np.exp(-5.0 * (x**2 + y**2))
# X-Y轴分为15*15的网格
y, x = np.mgrid[-1:1:15j, -1:1:15j] #❷
fvals = func(x, y) # 计算每个网格点上的函数值
# 二维插值
newfunc = interpolate.interp2d(x, y, fvals, kind='cubic') #❸
# 计算100*100的网格上的插值
xnew = np.linspace(-1, 1, 100)
ynew = np.linspace(-1, 1, 100)
fnew = newfunc(xnew, ynew) #❹
#%hide
pl.subplot(121)
pl.imshow(
fvals,
extent=[-1, 1, -1, 1],
cmap=pl.cm.jet,
interpolation='nearest',
origin="lower")
pl.title("fvals")
pl.subplot(122)
pl.imshow(
fnew,
extent=[-1, 1, -1, 1],
cmap=pl.cm.jet,
interpolation='nearest',
origin="lower")
pl.title("fnew")
pl.show()
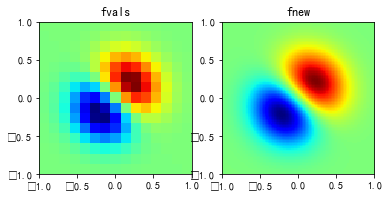
griddata
WARNING
griddata()使用欧几里得距离计算插值。如果 K 维空间中每个维度的取值范围相差较大,则应先将数据正规化,然后使用griddata()进行插值运算。
#%fig=使用gridata进行二维插值
# 计算随机N个点的坐标,以及这些点对应的函数值
N = 200
np.random.seed(42)
x = np.random.uniform(-1, 1, N)
y = np.random.uniform(-1, 1, N)
z = func(x, y)
yg, xg = np.mgrid[-1:1:100j, -1:1:100j]
xi = np.c_[xg.ravel(), yg.ravel()]
methods = 'nearest', 'linear', 'cubic'
zgs = [
interpolate.griddata((x, y), z, xi, method=method).reshape(100, 100)
for method in methods
]
#%hide
fig, axes = pl.subplots(1, 3, figsize=(11.5, 3.5))
for ax, method, zg in zip(axes, methods, zgs):
ax.imshow(
zg,
extent=[-1, 1, -1, 1],
cmap=pl.cm.jet,
interpolation='nearest',
origin="lower")
ax.set_xlabel(method)
ax.scatter(x, y, c=z)

径向基函数插值
#%fig=一维RBF插值
from scipy.interpolate import Rbf
x1 = np.array([-1, 0, 2.0, 1.0])
y1 = np.array([1.0, 0.3, -0.5, 0.8])
funcs = ['multiquadric', 'gaussian', 'linear']
nx = np.linspace(-3, 4, 100)
rbfs = [Rbf(x1, y1, function=fname) for fname in funcs] #❶
rbf_ys = [rbf(nx) for rbf in rbfs] #❷
#%hide
pl.plot(x1, y1, "o")
for fname, ny in zip(funcs, rbf_ys):
pl.plot(nx, ny, label=fname, lw=2)
pl.ylim(-1.0, 1.5)
pl.legend()
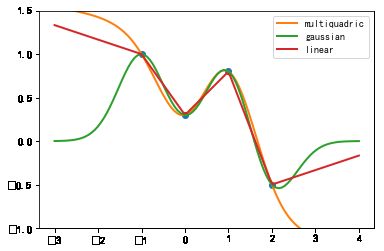 output_20_1
output_20_1
for fname, rbf in zip(funcs, rbfs):
print (fname, rbf.nodes)
multiquadric [-0.88822885 2.17654513 1.42877511 -2.67919021]
gaussian [ 1.00321945 -0.02345964 -0.65441716 0.91375159]
linear [-0.26666667 0.6 0.73333333 -0.9 ]
#%fig=二维径向基函数插值
rbfs = [Rbf(x, y, z, function=fname) for fname in funcs]
rbf_zg = [rbf(xg, yg).reshape(xg.shape) for rbf in rbfs]
#%hide
fig, axes = pl.subplots(1, 3, figsize=(11.5, 3.5))
for ax, fname, zg in zip(axes, funcs, rbf_zg):
ax.imshow(
zg,
extent=[-1, 1, -1, 1],
cmap=pl.cm.jet,
interpolation='nearest',
origin="lower")
ax.set_xlabel(fname)
ax.scatter(x, y, c=z)

#%fig=`epsilon`参数指定径向基函数中数据点的作用范围
epsilons = 0.1, 0.15, 0.3
rbfs = [Rbf(x, y, z, function="gaussian", epsilon=eps) for eps in epsilons]
zgs = [rbf(xg, yg).reshape(xg.shape) for rbf in rbfs]
#%hide
fig, axes = pl.subplots(1, 3, figsize=(11.5, 3.5))
for ax, eps, zg in zip(axes, epsilons, zgs):
ax.imshow(
zg,
extent=[-1, 1, -1, 1],
cmap=pl.cm.jet,
interpolation='nearest',
origin="lower")
ax.set_xlabel("eps=%g" % eps)
ax.scatter(x, y, c=z)

8.稀疏矩阵-sparse
%matplotlib inline
import numpy as np
import pylab as pl
from scipy import sparse
from scipy.sparse import csgraph
稀疏矩阵的储存形式
from scipy import sparse
a = sparse.dok_matrix((10, 5))
a[2, 3] = 1.0
a[3, 3] = 2.0
a[4, 3] = 3.0
print(a.keys())
print(a.values())
dict_keys([(2, 3), (3, 3), (4, 3)])
dict_values([1.0, 2.0, 3.0])
b = sparse.lil_matrix((10, 5))
b[2, 3] = 1.0
b[3, 4] = 2.0
b[3, 2] = 3.0
print(b.data)
print(b.rows)
[list([]) list([]) list([1.0]) list([3.0, 2.0]) list([]) list([]) list([])
list([]) list([]) list([])]
[list([]) list([]) list([3]) list([2, 4]) list([]) list([]) list([])
list([]) list([]) list([])]
row = [2, 3, 3, 2]
col = [3, 4, 2, 3]
data = [1, 2, 3, 10]
c = sparse.coo_matrix((data, (row, col)), shape=(5, 6))
print (c.col, c.row, c.data)
print (c.toarray())
[3 4 2 3] [2 3 3 2] [ 1 2 3 10]
[[ 0 0 0 0 0 0]
[ 0 0 0 0 0 0]
[ 0 0 0 11 0 0]
[ 0 0 3 0 2 0]
[ 0 0 0 0 0 0]]
矩阵向量相乘
import numpy as np
from scipy.sparse import csr_matrix
A = csr_matrix([[1, 2, 0], [0, 0, 3], [4, 0, 5]])
v = np.array([1, 0, -1])
A.dot(v)
array([ 1, -3, -1], dtype=int32)
示例1
构造一个1000x1000 lil_matrix并添加值:
from scipy.sparse import lil_matrix
from scipy.sparse.linalg import spsolve
from numpy.linalg import solve, norm
from numpy.random import rand
A = lil_matrix((1000, 1000))
A[0, :100] = rand(100)
A[1, 100:200] = A[0, :100]
A.setdiag(rand(1000))
现在将其转换为CSR格式,并求解 的 :
A = A.tocsr()
b = rand(1000)
x = spsolve(A, b)
将其转换为密集矩阵并求解,并检查结果是否相同:
x_ = solve(A.toarray(), b)
现在我们可以使用以下公式计算错误的范数:
err = norm(x-x_)
err < 1e-10
True
示例2
构造COO格式的矩阵:
from scipy import sparse
from numpy import array
I = array([0,3,1,0])
J = array([0,3,1,2])
V = array([4,5,7,9])
A = sparse.coo_matrix((V,(I,J)),shape=(4,4))
注意,索引不需要排序。
转换为CSR或CSC时,将对重复的(i,j)条目进行求和。
I = array([0,0,1,3,1,0,0])
J = array([0,2,1,3,1,0,0])
V = array([1,1,1,1,1,1,1])
B = sparse.coo_matrix((V,(I,J)),shape=(4,4)).tocsr()
这对于构造有限元刚度矩阵和质量矩阵是有用的。
9.图像处理-ndimage
import numpy as np
import pylab as pl
形态学图像处理
import numpy as np
def expand_image(img, value, out=None, size = 10):
if out is None:
w, h = img.shape
out = np.zeros((w*size, h*size),dtype=np.uint8)
tmp = np.repeat(np.repeat(img,size,0),size,1)
out[:,:] = np.where(tmp, value, out)
out[::size,:] = 0
out[:,::size] = 0
return out
def show_image(*imgs):
for idx, img in enumerate(imgs, 1):
ax = pl.subplot(1, len(imgs), idx)
pl.imshow(img, cmap="gray")
ax.set_axis_off()
pl.subplots_adjust(0.02, 0, 0.98, 1, 0.02, 0)
膨胀和腐蚀
#%fig=四连通和八连通的膨胀运算
from scipy.ndimage import morphology
def dilation_demo(a, structure=None):
b = morphology.binary_dilation(a, structure)
img = expand_image(a, 255)
return expand_image(np.logical_xor(a,b), 150, out=img)
a = pl.imread("scipy_morphology_demo.png")[:,:,0].astype(np.uint8)
img1 = expand_image(a, 255)
img2 = dilation_demo(a)
img3 = dilation_demo(a, [[1,1,1],[1,1,1],[1,1,1]])
show_image(img1, img2, img3)

#%fig=不同结构元素的膨胀效果
img4 = dilation_demo(a, [[0,0,0],[1,1,1],[0,0,0]])
img5 = dilation_demo(a, [[0,1,0],[0,1,0],[0,1,0]])
img6 = dilation_demo(a, [[0,1,0],[0,1,0],[0,0,0]])
show_image(img4, img5, img6)

#%fig=四连通和八连通的腐蚀运算
def erosion_demo(a, structure=None):
b = morphology.binary_erosion(a, structure)
img = expand_image(a, 255)
return expand_image(np.logical_xor(a,b), 100, out=img)
img1 = expand_image(a, 255)
img2 = erosion_demo(a)
img3 = erosion_demo(a, [[1,1,1],[1,1,1],[1,1,1]])
show_image(img1, img2, img3)

Hit和Miss
#%fig=Hit和Miss运算
def hitmiss_demo(a, structure1, structure2):
b = morphology.binary_hit_or_miss(a, structure1, structure2)
img = expand_image(a, 100)
return expand_image(b, 255, out=img)
img1 = expand_image(a, 255)
img2 = hitmiss_demo(a, [[0,0,0],[0,1,0],[1,1,1]], [[1,0,0],[0,0,0],[0,0,0]])
img3 = hitmiss_demo(a, [[0,0,0],[0,0,0],[1,1,1]], [[1,0,0],[0,1,0],[0,0,0]])
show_image(img1, img2, img3)

#%fig=使用Hit和Miss进行细线化运算
def skeletonize(img):
h1 = np.array([[0, 0, 0],[0, 1, 0],[1, 1, 1]]) #❶
m1 = np.array([[1, 1, 1],[0, 0, 0],[0, 0, 0]])
h2 = np.array([[0, 0, 0],[1, 1, 0],[0, 1, 0]])
m2 = np.array([[0, 1, 1],[0, 0, 1],[0, 0, 0]])
hit_list = []
miss_list = []
for k in range(4): #❷
hit_list.append(np.rot90(h1, k))
hit_list.append(np.rot90(h2, k))
miss_list.append(np.rot90(m1, k))
miss_list.append(np.rot90(m2, k))
img = img.copy()
while True:
last = img
for hit, miss in zip(hit_list, miss_list):
hm = morphology.binary_hit_or_miss(img, hit, miss) #❸
# 从图像中删除hit_or_miss所得到的白色点
img = np.logical_and(img, np.logical_not(hm)) #❹
# 如果处理之后的图像和处理前的图像相同,则结束处理
if np.all(img == last): #❺
break
return img
a = pl.imread("scipy_morphology_demo2.png")[:,:,0].astype(np.uint8)
b = skeletonize(a)
#%hide
_, (ax1, ax2) = pl.subplots(1, 2, figsize=(9, 3))
ax1.imshow(a, cmap="gray", interpolation="nearest")
ax2.imshow(b, cmap="gray", interpolation="nearest")
ax1.set_axis_off()
ax2.set_axis_off()
pl.subplots_adjust(0.02, 0, 0.98, 1, 0.02, 0)

图像分割
squares = pl.imread("suqares.jpg")
squares = (squares[:,:,0] < 200).astype(np.uint8)
from scipy.ndimage import morphology
squares_dt = morphology.distance_transform_cdt(squares)
print ("各种距离值", np.unique(squares_dt))
各种距离值 [ 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23
24 25 26 27]
squares_core = (squares_dt > 8).astype(np.uint8)
from scipy.ndimage.measurements import label, center_of_mass
def random_palette(labels, count, seed=1):
np.random.seed(seed)
palette = np.random.rand(count+1, 3)
palette[0,:] = 0
return palette[labels]
labels, count = label(squares_core)
h, w = labels.shape
centers = np.array(center_of_mass(labels, labels, index=range(1, count+1)), np.int)
cores = random_palette(labels, count)
index = morphology.distance_transform_cdt(1-squares_core,
return_distances=False,
return_indices=True) #❶
near_labels = labels[index[0], index[1]] #❷
mask = (squares - squares_core).astype(bool)
labels2 = labels.copy()
labels2[mask] = near_labels[mask] #❸
separated = random_palette(labels2, count)
#%figonly=矩形区域分割算法各个步骤的输出图像
fig, axes = pl.subplots(2, 3, figsize=(7.5, 5.0), )
fig.delaxes(axes[1, 2])
axes[0, 0].imshow(squares, cmap="gray");
axes[0, 1].imshow(squares_dt)
axes[0, 2].imshow(squares_core, cmap="gray")
ax = axes[1, 0]
ax.imshow(cores)
center_y, center_x = centers.T
ax.plot(center_x, center_y, "o", color="white")
ax.set_xlim(0, w)
ax.set_ylim(h, 0)
axes[1, 1].imshow(separated)
for ax in axes.ravel():
ax.axis("off")
fig.subplots_adjust(wspace=0.01, hspace=0.01)

10.空间算法库-spatial
import numpy as np
import pylab as pl
from scipy import spatial
import matplotlib as mpl
mpl.rcParams['font.sans-serif'] = ['SimHei']
计算最近旁点
x = np.sort(np.random.rand(100))
idx = np.searchsorted(x, 0.5)
print (x[idx], x[idx - 1]) #距离0.5最近的数是这两个数中的一个
0.5244435681885733 0.4982156075770372
from scipy import spatial
np.random.seed(42)
N = 100
points = np.random.uniform(-1, 1, (N, 2))
kd = spatial.cKDTree(points)
targets = np.array([(0, 0), (0.5, 0.5), (-0.5, 0.5), (0.5, -0.5), (-0.5, -0.5)])
dist, idx = kd.query(targets, 3)
r = 0.2
idx2 = kd.query_ball_point(targets, r)
idx2
array([list([48]), list([37, 78]), list([22, 79, 92]), list([6, 35, 58]),
list([7, 42, 55, 83])], dtype=object)
idx3 = kd.query_pairs(0.1) - kd.query_pairs(0.08)
idx3
{(1, 46),
(3, 21),
(3, 82),
(3, 95),
(5, 16),
(9, 30),
(10, 87),
(11, 42),
(11, 97),
(18, 41),
(29, 74),
(32, 51),
(37, 78),
(39, 61),
(41, 61),
(50, 84),
(55, 83),
(73, 81)}
#%figonly=用cKDTree寻找近旁点
x, y = points.T
colors = "r", "b", "g", "y", "k"
fig, (ax1, ax2, ax3) = pl.subplots(1, 3, figsize=(12, 4))
for ax in ax1, ax2, ax3:
ax.set_aspect("equal")
ax.plot(x, y, "o", markersize=4)
for ax in ax1, ax2:
for i in range(len(targets)):
c = colors[i]
tx, ty = targets[i]
ax.plot([tx], [ty], "*", markersize=10, color=c)
for i in range(len(targets)):
nx, ny = points[idx[i]].T
ax1.plot(nx, ny, "o", markersize=10, markerfacecolor="None",
markeredgecolor=colors[i], markeredgewidth=1)
nx, ny = points[idx2[i]].T
ax2.plot(nx, ny, "o", markersize=10, markerfacecolor="None",
markeredgecolor=colors[i], markeredgewidth=1)
ax2.add_artist(pl.Circle(targets[i], r, fill=None, linestyle="dashed"))
for pidx1, pidx2 in idx3:
sx, sy = points[pidx1]
ex, ey = points[pidx2]
ax3.plot([sx, ex], [sy, ey], "r", linewidth=2, alpha=0.6)
ax1.set_xlabel(u"搜索最近的3个近旁点")
ax2.set_xlabel(u"搜索距离在0.2之内的所有近旁点")
ax3.set_xlabel(u"搜索所有距离在0.08到0.1之间的点对");

from scipy.spatial import distance
dist1 = distance.squareform(distance.pdist(points))
dist2 = distance.cdist(points, targets)
print(dist1.shape)
print(dist2.shape)
(100, 100)
(100, 5)
print (dist[:, 0]) # cKDTree.query()返回的与targets最近的距离
print (np.min(dist2, axis=0))
[0.15188266 0.09595807 0.05009422 0.11180181 0.19015485]
[0.15188266 0.09595807 0.05009422 0.11180181 0.19015485]
dist1[np.diag_indices(len(points))] = np.inf
nearest_pair = np.unravel_index(np.argmin(dist1), dist1.shape)
print (nearest_pair, dist1[nearest_pair])
(22, 92) 0.005346210248158245
dist, idx = kd.query(points, 2)
print (idx[np.argmin(dist[:, 1])], np.min(dist[:, 1]))
[22 92] 0.005346210248158245
N = 1000000
start = np.random.uniform(0, 100, N)
span = np.random.uniform(0.01, 1, N)
span = np.clip(span, 2, 100)
end = start + span
def naive_count_at(start, end, time):
mask = (start < time) & (end > time)
return np.sum(mask)
#%figonly=使用二维K-d树搜索指定区间的在线用户
def _():
N = 100
start = np.random.uniform(0, 100, N)
span = np.random.normal(40, 10, N)
span = np.clip(span, 2, 100)
end = start + span
time = 40
fig, ax = pl.subplots(figsize=(8, 6))
ax.scatter(start, end)
mask = (start < time) & (end > time)
start2, end2 = start[mask], end[mask]
ax.scatter(start2, end2, marker="x", color="red")
rect = pl.Rectangle((-20, 40), 60, 120, alpha=0.3)
ax.add_patch(rect)
ax.axhline(time, color="k", ls="--")
ax.axvline(time, color="k", ls="--")
ax.set_xlabel("Start")
ax.set_ylabel("End")
ax.set_xlim(-20, 120)
ax.set_ylim(-20, 160)
ax.plot([0, 120], [0, 120])
_()

class KdSearch(object):
def __init__(self, start, end, leafsize=10):
self.tree = spatial.cKDTree(np.c_[start, end], leafsize=leafsize)
self.max_time = np.max(end)
def count_at(self, time):
max_time = self.max_time
to_search = spatial.cKDTree([[time - max_time, time + max_time]])
return self.tree.count_neighbors(to_search, max_time, p=np.inf)
naive_count_at(start, end, 40) == KdSearch(start, end).count_at(40)
True
QUESTION
请读者研究点数
N和leafsize参数与创建 K-d 树和搜索时间之间的关系。
凸包
np.random.seed(42)
points2d = np.random.rand(10, 2)
ch2d = spatial.ConvexHull(points2d)
print(ch2d.simplices)
print(ch2d.vertices)
[[2 5]
[2 6]
[0 5]
[1 6]
[1 0]]
[5 2 6 1 0]
#%fig=二维平面上的凸包
poly = pl.Polygon(points2d[ch2d.vertices], fill=None, lw=2, color="r", alpha=0.5)
ax = pl.subplot(aspect="equal")
pl.plot(points2d[:, 0], points2d[:, 1], "go")
for i, pos in enumerate(points2d):
pl.text(pos[0], pos[1], str(i), color="blue")
ax.add_artist(poly);
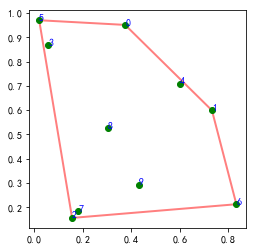
np.random.seed(42)
points3d = np.random.rand(40, 3)
ch3d = spatial.ConvexHull(points3d)
ch3d.simplices.shape

(38, 3)
沃罗诺伊图
points2d = np.array([[0.2, 0.1], [0.5, 0.5], [0.8, 0.1],
[0.5, 0.8], [0.3, 0.6], [0.7, 0.6], [0.5, 0.35]])
vo = spatial.Voronoi(points2d)
print(vo.vertices); print(vo.regions); print(vo.ridge_vertices)
[[0.5 0.045 ]
[0.245 0.351 ]
[0.755 0.351 ]
[0.3375 0.425 ]
[0.6625 0.425 ]
[0.45 0.65 ]
[0.55 0.65 ]]
[[-1, 0, 1], [-1, 0, 2], [], [6, 4, 3, 5], [5, -1, 1, 3], [4, 2, 0, 1, 3], [6, -1, 2, 4], [6, -1, 5]]
[[-1, 0], [0, 1], [-1, 1], [0, 2], [-1, 2], [3, 5], [3, 4], [4, 6], [5, 6], [1, 3], [-1, 5], [2, 4], [-1, 6]]
bound = np.array([[-100, -100], [-100, 100],
[ 100, 100], [ 100, -100]])
vo2 = spatial.Voronoi(np.vstack((points2d, bound)))
#%figonly=沃罗诺伊图将空间分割为多个区域
fig, (ax1, ax2) = pl.subplots(1, 2, figsize=(9, 4.5))
ax1.set_aspect("equal")
ax2.set_aspect("equal")
spatial.voronoi_plot_2d(vo, ax=ax1)
for i, v in enumerate(vo.vertices):
ax1.text(v[0], v[1], str(i), color="red")
for i, p in enumerate(points2d):
ax1.text(p[0], p[1], str(i), color="blue")
n = len(points2d)
color = pl.cm.rainbow(np.linspace(0, 1, n))
for i in range(n):
idx = vo2.point_region[i]
region = vo2.regions[idx]
poly = pl.Polygon(vo2.vertices[region], facecolor=color[i], alpha=0.5, zorder=0)
ax2.add_artist(poly)
ax2.scatter(points2d[:, 0], points2d[:, 1], s=40, c=color, linewidths=2, edgecolors="k")
ax2.plot(vo2.vertices[:, 0], vo2.vertices[:, 1], "ro", ms=6)
for ax in ax1, ax2:
ax.set_xlim(0, 1)
ax.set_ylim(0, 1)
 output_26_1
output_26_1
print (vo.point_region)
print (vo.regions[6])
[0 3 1 7 4 6 5]
[6, -1, 2, 4]
德劳内三角化
x = np.array([46.445, 263.251, 174.176, 280.899, 280.899,
189.358, 135.521, 29.638, 101.907, 226.665])
y = np.array([287.865, 250.891, 287.865, 160.975, 54.252,
160.975, 232.404, 179.187, 35.765, 71.361])
points2d = np.c_[x, y]
dy = spatial.Delaunay(points2d)
vo = spatial.Voronoi(points2d)
print(dy.simplices)
print(vo.vertices)
[[8 5 7]
[1 5 3]
[5 6 7]
[6 0 7]
[0 6 2]
[6 1 2]
[1 6 5]
[9 5 8]
[4 9 8]
[5 9 3]
[9 4 3]]
[[104.58977484 127.03566055]
[235.1285 198.68143374]
[107.83960707 155.53682482]
[ 71.22104881 228.39479887]
[110.3105 291.17642838]
[201.40695449 227.68436282]
[201.61895891 226.21958623]
[152.96231864 93.25060083]
[205.40381294 -90.5480267 ]
[235.1285 127.45701644]
[267.91709907 107.6135 ]]
#%fig=德劳内三角形的外接圆与圆心
cx, cy = vo.vertices.T
ax = pl.subplot(aspect="equal")
spatial.delaunay_plot_2d(dy, ax=ax)
ax.plot(cx, cy, "r.")
for i, (cx, cy) in enumerate(vo.vertices):
px, py = points2d[dy.simplices[i, 0]]
radius = np.hypot(cx - px, cy - py)
circle = pl.Circle((cx, cy), radius, fill=False, ls="dotted")
ax.add_artist(circle)
ax.set_xlim(0, 300)
ax.set_ylim(0, 300);

![]()
往期精彩回顾
那些年做的学术公益-你不是一个人在战斗适合初学者入门人工智能的路线及资料下载机器学习在线手册深度学习在线手册备注:加入本站微信群或者qq群,请回复“加群”加入知识星球(4500+用户,ID:92416895),请回复“知识星球”