吴恩达Coursera深度学习课程 DeepLearning.ai 编程作业——Regularization(2-1.2)
如果数据集没有很大,同时在训练集上又拟合得很好,但是在测试集的效果却不是很好,这时候就要使用正则化来使得其拟合能力不会那么强。
import numpy as np
import sklearn
import matplotlib.pyplot as plt
import sklearn.datasets
from reg_utils import load_2D_dataset,compute_cost,forward_propagation,predict,predict_dec,relu,sigmoid
from reg_utils import initialize_parameters,backward_propagation,update_parameters,plot_decision_boundary
import scipy.io
plt.rcParams["figure.figsize"]=(7,4) #the figure size,which wight is 7,while height is 4
plt.rcParams["image.interpolation"]="nearest" #
plt.rcParams["image.cmap"]="gray"
train_x,train_y,test_x,test_x,test_y=load_2D_dataset()
import numpy as np
import matplotlib.pyplot as plt
import h5py
import sklearn
import sklearn.datasets
import sklearn.linear_model
import scipy.io
####reg_utils.py##### 以下是reg_utils.py 文件的内容
def sigmoid(x):
s = 1/(1+np.exp(-x))
return s
def relu(x):
s = np.maximum(0,x)
return s
def load_planar_dataset(seed):
np.random.seed(seed)
m = 400 # number of examples
N = int(m/2) # number of points per class
D = 2 # dimensionality
X = np.zeros((m,D)) # data matrix where each row is a single example
Y = np.zeros((m,1), dtype='uint8') # labels vector (0 for red, 1 for blue)
a = 4 # maximum ray of the flower
for j in range(2):
ix = range(N*j,N*(j+1))
t = np.linspace(j*3.12,(j+1)*3.12,N) + np.random.randn(N)*0.2 # theta
r = a*np.sin(4*t) + np.random.randn(N)*0.2 # radius
X[ix] = np.c_[r*np.sin(t), r*np.cos(t)]
Y[ix] = j
X = X.T
Y = Y.T
return X, Y
def initialize_parameters(layer_dims):
"""
Arguments:
layer_dims -- python array (list) containing the dimensions of each layer in our network
Returns:
parameters -- python dictionary containing your parameters "W1", "b1", ..., "WL", "bL":
W1 -- weight matrix of shape (layer_dims[l], layer_dims[l-1])
b1 -- bias vector of shape (layer_dims[l], 1)
Wl -- weight matrix of shape (layer_dims[l-1], layer_dims[l])
bl -- bias vector of shape (1, layer_dims[l])
Tips:
- For example: the layer_dims for the "Planar Data classification model" would have been [2,2,1].
This means W1's shape was (2,2), b1 was (1,2), W2 was (2,1) and b2 was (1,1). Now you have to generalize it!
- In the for loop, use parameters['W' + str(l)] to access Wl, where l is the iterative integer.
"""
np.random.seed(3)
parameters = {}
L = len(layer_dims) # number of layers in the network
for l in range(1, L):
parameters['W' + str(l)] = np.random.randn(layer_dims[l], layer_dims[l-1]) / np.sqrt(layer_dims[l-1])
parameters['b' + str(l)] = np.zeros((layer_dims[l], 1))
assert(parameters['W' + str(l)].shape == layer_dims[l], layer_dims[l-1])
assert(parameters['W' + str(l)].shape == layer_dims[l], 1)
return parameters
def forward_propagation(X, parameters):
"""
Implements the forward propagation (and computes the loss) presented in Figure 2.
Arguments:
X -- input dataset, of shape (input size, number of examples)
parameters -- python dictionary containing your parameters "W1", "b1", "W2", "b2", "W3", "b3":
W1 -- weight matrix of shape ()
b1 -- bias vector of shape ()
W2 -- weight matrix of shape ()
b2 -- bias vector of shape ()
W3 -- weight matrix of shape ()
b3 -- bias vector of shape ()
Returns:
loss -- the loss function (vanilla logistic loss)
"""
# retrieve parameters
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
W3 = parameters["W3"]
b3 = parameters["b3"]
# LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SIGMOID
Z1 = np.dot(W1, X) + b1
A1 = relu(Z1)
Z2 = np.dot(W2, A1) + b2
A2 = relu(Z2)
Z3 = np.dot(W3, A2) + b3
A3 = sigmoid(Z3)
cache = (Z1, A1, W1, b1, Z2, A2, W2, b2, Z3, A3, W3, b3)
return A3, cache
def backward_propagation(X, Y, cache):
"""
Implement the backward propagation presented in figure 2.
Arguments:
X -- input dataset, of shape (input size, number of examples)
Y -- true "label" vector (containing 0 if cat, 1 if non-cat)
cache -- cache output from forward_propagation()
Returns:
gradients -- A dictionary with the gradients with respect to each parameter, activation and pre-activation variables
"""
m = X.shape[1]
(Z1, A1, W1, b1, Z2, A2, W2, b2, Z3, A3, W3, b3) = cache
dZ3 = A3 - Y
dW3 = 1./m * np.dot(dZ3, A2.T)
db3 = 1./m * np.sum(dZ3, axis=1, keepdims = True)
dA2 = np.dot(W3.T, dZ3)
dZ2 = np.multiply(dA2, np.int64(A2 > 0))
dW2 = 1./m * np.dot(dZ2, A1.T)
db2 = 1./m * np.sum(dZ2, axis=1, keepdims = True)
dA1 = np.dot(W2.T, dZ2)
dZ1 = np.multiply(dA1, np.int64(A1 > 0))
dW1 = 1./m * np.dot(dZ1, X.T)
db1 = 1./m * np.sum(dZ1, axis=1, keepdims = True)
gradients = {"dZ3": dZ3, "dW3": dW3, "db3": db3,
"dA2": dA2, "dZ2": dZ2, "dW2": dW2, "db2": db2,
"dA1": dA1, "dZ1": dZ1, "dW1": dW1, "db1": db1}
return gradients
def update_parameters(parameters, grads, learning_rate):
"""
Update parameters using gradient descent
Arguments:
parameters -- python dictionary containing your parameters:
parameters['W' + str(i)] = Wi
parameters['b' + str(i)] = bi
grads -- python dictionary containing your gradients for each parameters:
grads['dW' + str(i)] = dWi
grads['db' + str(i)] = dbi
learning_rate -- the learning rate, scalar.
Returns:
parameters -- python dictionary containing your updated parameters
"""
n = len(parameters) // 2 # number of layers in the neural networks
# Update rule for each parameter
for k in range(n):
parameters["W" + str(k+1)] = parameters["W" + str(k+1)] - learning_rate * grads["dW" + str(k+1)]
parameters["b" + str(k+1)] = parameters["b" + str(k+1)] - learning_rate * grads["db" + str(k+1)]
return parameters
def predict(X, y, parameters):
"""
This function is used to predict the results of a n-layer neural network.
Arguments:
X -- data set of examples you would like to label
parameters -- parameters of the trained model
Returns:
p -- predictions for the given dataset X
"""
m = X.shape[1]
p = np.zeros((1,m), dtype = np.int)
# Forward propagation
a3, caches = forward_propagation(X, parameters)
# convert probas to 0/1 predictions
for i in range(0, a3.shape[1]):
if a3[0,i] > 0.5:
p[0,i] = 1
else:
p[0,i] = 0
# print results
#print ("predictions: " + str(p[0,:]))
#print ("true labels: " + str(y[0,:]))
print("Accuracy: " + str(np.mean((p[0,:] == y[0,:]))))
return p
def compute_cost(a3, Y):
"""
Implement the cost function
Arguments:
a3 -- post-activation, output of forward propagation
Y -- "true" labels vector, same shape as a3
Returns:
cost - value of the cost function
"""
m = Y.shape[1]
logprobs = np.multiply(-np.log(a3),Y) + np.multiply(-np.log(1 - a3), 1 - Y)
cost = 1./m * np.nansum(logprobs)
return cost
def load_dataset():
train_dataset = h5py.File('datasets/train_catvnoncat.h5', "r")
train_set_x_orig = np.array(train_dataset["train_set_x"][:]) # your train set features
train_set_y_orig = np.array(train_dataset["train_set_y"][:]) # your train set labels
test_dataset = h5py.File('datasets/test_catvnoncat.h5', "r")
test_set_x_orig = np.array(test_dataset["test_set_x"][:]) # your test set features
test_set_y_orig = np.array(test_dataset["test_set_y"][:]) # your test set labels
classes = np.array(test_dataset["list_classes"][:]) # the list of classes
train_set_y = train_set_y_orig.reshape((1, train_set_y_orig.shape[0]))
test_set_y = test_set_y_orig.reshape((1, test_set_y_orig.shape[0]))
train_set_x_orig = train_set_x_orig.reshape(train_set_x_orig.shape[0], -1).T
test_set_x_orig = test_set_x_orig.reshape(test_set_x_orig.shape[0], -1).T
train_set_x = train_set_x_orig/255
test_set_x = test_set_x_orig/255
return train_set_x, train_set_y, test_set_x, test_set_y, classes
def predict_dec(parameters, X):
"""
Used for plotting decision boundary.
Arguments:
parameters -- python dictionary containing your parameters
X -- input data of size (m, K)
Returns
predictions -- vector of predictions of our model (red: 0 / blue: 1)
"""
# Predict using forward propagation and a classification threshold of 0.5
a3, cache = forward_propagation(X, parameters)
predictions = (a3>0.5)
return predictions
def load_planar_dataset(randomness, seed):
np.random.seed(seed)
m = 50
N = int(m/2) # number of points per class
D = 2 # dimensionality
X = np.zeros((m,D)) # data matrix where each row is a single example
Y = np.zeros((m,1), dtype='uint8') # labels vector (0 for red, 1 for blue)
a = 2 # maximum ray of the flower
for j in range(2):
ix = range(N*j,N*(j+1))
if j == 0:
t = np.linspace(j, 4*3.1415*(j+1),N) #+ np.random.randn(N)*randomness # theta
r = 0.3*np.square(t) + np.random.randn(N)*randomness # radius
if j == 1:
t = np.linspace(j, 2*3.1415*(j+1),N) #+ np.random.randn(N)*randomness # theta
r = 0.2*np.square(t) + np.random.randn(N)*randomness # radius
X[ix] = np.c_[r*np.cos(t), r*np.sin(t)]
Y[ix] = j
X = X.T
Y = Y.T
return X, Y
def plot_decision_boundary(model, X, y):
# Set min and max values and give it some padding
x_min, x_max = X[0, :].min() - 1, X[0, :].max() + 1
y_min, y_max = X[1, :].min() - 1, X[1, :].max() + 1
h = 0.01
# Generate a grid of points with distance h between them
xx, yy = np.meshgrid(np.arange(x_min, x_max, h), np.arange(y_min, y_max, h))
# Predict the function value for the whole grid
print ("the input shape"+str(np.c_[xx.ravel(),yy.ravel()].shape))
Z = model(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
# Plot the contour and training examples
plt.contourf(xx, yy, Z, cmap=plt.cm.Spectral)
plt.ylabel('x2')
plt.xlabel('x1')
plt.scatter(X[0, :], X[1, :], c=y, cmap=plt.cm.Spectral)
plt.show()
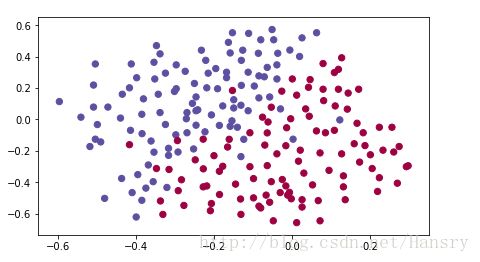
def load_2D_dataset():
data = scipy.io.loadmat('/home/hansry/python/DL/2-1/week1_Regularization/datasets/data.mat.mat')
train_X = data['X'].T
train_Y = data['y'].T
test_X = data['Xval'].T
test_Y = data['yval'].T
plt.scatter(train_X[0, :], train_X[1, :], c=train_Y, s=40, cmap=plt.cm.Spectral);
plt.show()
return train_X, train_Y, test_X, test_Y
- Non-regularization model
def model(X,Y,learning_rate=0.3,num_iterations=30000,print_cost=True,lambd=0,keep_prob=1):
grads={}
costs=[]
m=X.shape[1]
layers_dims=[X.shape[0],20,3,1]
#Initialize=initialize_parameters(layers_dims)
parameters=initialize_parameters(layers_dims)
for i in range(0, num_iterations):
# Forward propagation: LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SIGMOID.
if keep_prob == 1:
a3, cache = forward_propagation(X, parameters)
elif keep_prob < 1: #当keep_prob<1时,使用drop_out,其中keep_prob between 0 and 1
a3, cache,cache_Dl = forward_propagation_with_dropout(X, parameters, keep_prob)
# Cost function
if lambd == 0:
cost = compute_cost(a3, Y)
else: #如果lambd不等于0,那么使用regulrization
cost = compute_cost_with_regularization(a3, Y, parameters, lambd)
# Backward propagation.
assert(lambd==0 or keep_prob==1) # it is possible to use both L2 regularization and dropout,
# but this assignment will only explore one at a time
if lambd == 0 and keep_prob == 1:
grads = backward_propagation(X, Y, cache)
elif lambd != 0:
grads = backward_propagation_with_regularization(X, Y, cache, lambd)
elif keep_prob < 1:
grads = backward_propogation_with_dropout(X,Y,cache,cache_Dl,keep_prob)
# Update parameters.
parameters = update_parameters(parameters, grads, learning_rate)
# Print the loss every 10000 iterations
if print_cost and i % 10000 == 0:
print("Cost after iteration {}: {}".format(i, cost))
if print_cost and i % 1000 == 0:
costs.append(cost)
# plot the cost
plt.plot(costs)
plt.ylabel('cost')
plt.xlabel('iterations (x1,000)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
return parameters
parameters=model(train_x,train_y,learning_rate=0.3,num_iterations=30000,print_cost=True,lambd=0,keep_prob=1)
print train_x.shape
print("On the train do not use regularization and dropout")
train_prediction=predict(train_x,train_y,parameters)
print ("On the test do not use regularization and dropout")
test_prediction=predict(test_x,test_y,parameters)
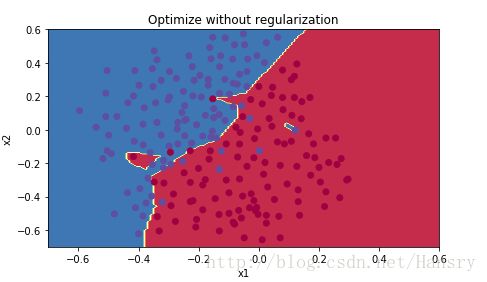
plt.title("Optimize without regularization")
axes=plt.gca()
axes.set_xlim([-0.7,0.6])
axes.set_ylim([-0.7,0.6])
plot_decision_boundary(lambda x: predict_dec(parameters,x.T),train_x,train_y)
Expected Output:
Cost after iteration 0: 0.655741252348
Cost after iteration 10000: 0.163299875257
Cost after iteration 20000: 0.138516424233
On the train do not use regularization and dropout
Accuracy: 0.947867298578
On the test do not use regularization and dropout
Accuracy: 0.915
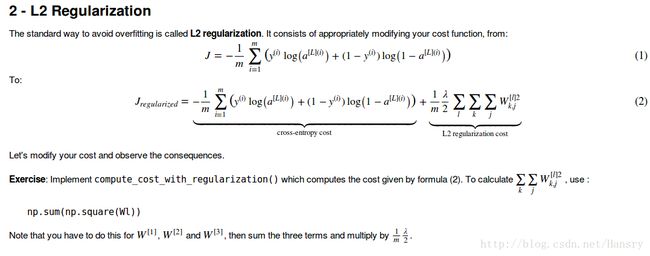
- L2 Regularization
def compute_cost_with_regularization(A3,Y,parameters,lamda): #将L2正则化添加到公式中
m=Y.shape[1]
L=len(parameters)//2
regularization_sum=0.0
for l in range(L):
regularization_sum += np.sum(np.square(parameters["W"+str(l+1)]))
regularization_cost=(lamda/(2.0*m))*regularization_sum
cost_entropy=compute_cost(A3,Y)
cost=cost_entropy+regularization_cost
return cost
'''
a3, Y_assess, parameters=compute_cost_with_regularization_test_case()
cost=compute_cost_with_regularization(a3,Y_assess,parameters,lamda=0.1)
print cost
'''
def backward_propagation_with_regularization(X,Y,caches,lamda): #这个函数对于多层神经网络均适用
L=len(caches)//4
cache={}
m=X.shape[1]
grads={}
for i in range(0,L):
cache["Z"+str(i+1)]=caches[i*4]
cache["A"+str(i+1)]=caches[i*4+1]
cache["W"+str(i+1)]=caches[i*4+2]
cache["b"+str(i+1)]=caches[i*4+3]
grads["dZ"+str(L)]=cache["A"+str(L)]-Y
grads["dW"+str(L)]=(1.0/m)*np.dot(grads["dZ"+str(L)],cache["A"+str(L-1)].T)+np.float(lamda/m)*cache["W"+str(L)]
grads["db"+str(L)]=(1.0/m)*np.sum(grads["dZ"+str(L)],axis=1,keepdims=True)
grads["dA"+str(L-1)]=np.dot(cache["W"+str(L)].T,grads["dZ"+str(L)])
for l in reversed(range(1,L)):
if l > 1:
grads["dZ"+str(l)]=np.multiply(grads["dA"+str(l)],np.int64(cache["A"+str(l)]>0))
# grads["dW"+str(l)]=cache["A"+str(L)]-Y
grads["dW"+str(l)]=(1.0/m)*np.dot(grads["dZ"+str(l)],cache["A"+str(l-1)].T)+np.float(lamda/m)*cache["W"+str(l)]
grads["db"+str(l)]=(1.0/m)*np.sum(grads["dZ"+str(l)],axis=1,keepdims=True)
grads["dA"+str(l-1)]=np.dot(cache["W"+str(l)].T,grads["dZ"+str(l)])
elif l==1:
grads["dZ"+str(1)]=np.multiply(grads["dA"+str(1)],np.int64(cache["A"+str(1)]>0))
grads["dW"+str(1)]=(1.0/m)*np.dot(grads["dZ"+str(1)],X.T)+np.float(lamda/m)*cache["W"+str(1)]
grads["db"+str(1)]=(1.0/m)*np.sum(grads["dZ"+str(1)],axis=1,keepdims=True)
return grads
#test the correctness of backward_propagation
"""
X_assess,Y_assess,cache=backward_propagation_with_regularization_test_case()
grads=backward_propagation_with_regularization(X_assess,Y_assess,cache,lamda=0.7)
print ("dW1 :"+str(grads["dW1"]))
print ("dW2 :"+str(grads["dW2"]))
print ("dW3 :"+str(grads["dW3"]))
"""
#remerber to recover the codes below
parameters_with_regularization=model(train_x,train_y,learning_rate=0.3,num_iterations=30000,print_cost=True,lambd=0.7,keep_prob=1)
print("On the train with regularization")
train_prediction_regularization=predict(train_x,train_y,parameters_with_regularization)
print ("On the test with regularization")
test_prediction_regularization=predict(test_x,test_y,parameters_with_regularization)
plt.title("Optimize with regularization")
axes=plt.gca()
axes.set_xlim([-0.7,0.6])
axes.set_ylim([-0.7,0.6])
plot_decision_boundary(lambda x: predict_dec(parameters_with_regularization,x.T),train_x,train_y)
Expected output:
Cost after iteration 0: 0.697448449313
Cost after iteration 10000: 0.268491887328
Cost after iteration 20000: 0.268091633713
On the train with regularization
Accuracy: 0.938388625592
On the test with regularization
Accuracy: 0.93
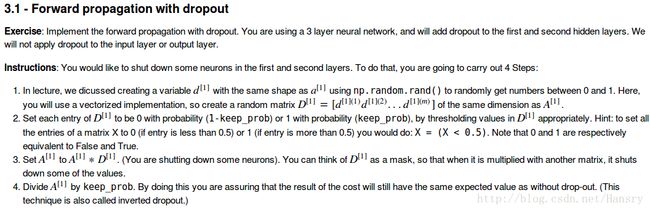
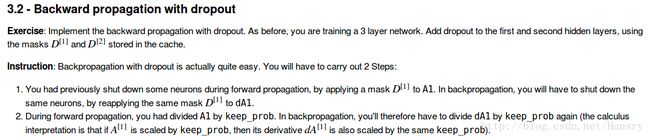
3.forward and backward propagation with dropout
def forward_propagation_with_dropout(X,parameters,keep_prob=0.5):
cache=[] #list
cache_D={} #dictionary
L=len(parameters)//2
W1=parameters["W1"]
b1=parameters["b1"]
Z1=np.dot(W1,X)+b1
A1=relu(Z1)
cache=[Z1,A1,W1,b1]
np.random.seed(1)
for l in range(1,L):
cache_D["D"+str(l)]=np.random.rand(cache[4*(l-1)+1].shape[0],cache[4*(l-1)+1].shape[1]) #randn(A1.shape[0],A1.shape[]1)
cache_D["D"+str(l)]=np.where(cache_D["D"+str(l)]<=keep_prob,1,0) # D1
cache[4*(l-1)+1]=np.multiply(cache[4*(l-1)+1],cache_D["D"+str(l)]) # A1,eliminate some neure randomly
cache[4*(l-1)+1]=cache[4*(l-1)+1]/keep_prob
cache_4l=np.dot(parameters["W"+str(l+1)],cache[4*(l-1)+1])+parameters["b"+str(l+1)] # cache[4*l] Z2
cache.append(cache_4l)
if l==(L-1):
cache_4l_plus1=sigmoid(cache[4*l]) #A2,A3,...
elif l0)) #dZ2(3,5)
if l>1:
grads["dW"+str(l)]=(1.0/m)*np.dot(grads["dZ"+str(l)],cache[4*(l-2)+1].T) # dW2(3,2)=dZ2(3,5)*A1(2,5).T
elif l==1:
grads["dW"+str(l)]=(1.0/m)*np.dot(grads["dZ"+str(l)],X.T)
grads["db"+str(l)]=(1.0/m)*np.sum(grads["dZ"+str(l)],axis=1,keepdims=True)
grads["dA"+str(l-1)]=np.dot(cache[4*(l-1)+2].T,grads["dZ"+str(l)])
return grads
parameters_with_dropout=model(train_x,train_y,keep_prob=0.86,learning_rate=0.3)
print ("On the train set with dropout:")
predictions_train = predict(train_x, train_y, parameters_with_dropout)
print ("On the test set with dropout:")
predictions_test = predict(test_x, test_y, parameters_with_dropout)
plt.title("the cost curve with ")
axes=plt.gca()
axes.set_xlim([-0.7,0.6])
axes.set_ylim([-0.7,0.6])
plot_decision_boundary(lambda x:predict_dec(parameters_with_dropout,x.T),train_x,train_y)
Expected output:
Cost after iteration 0: 0.654391240515
reg_utils.py:238: RuntimeWarning: divide by zero encountered in log
logprobs = np.multiply(-np.log(a3),Y) + np.multiply(-np.log(1 - a3), 1 - Y)
reg_utils.py:238: RuntimeWarning: invalid value encountered in multiply
logprobs = np.multiply(-np.log(a3),Y) + np.multiply(-np.log(1 - a3), 1 - Y)
Cost after iteration 10000: 0.0610169865749
Cost after iteration 20000: 0.0605824357985
On the train set with dropout:
Accuracy: 0.928909952607
On the test set with dropout:
Accuracy: 0.95