MATLAB机器学习决策树网格法优化参数学习笔记
适用于2017b版本以后的版本,之前的老版本可能有所出入
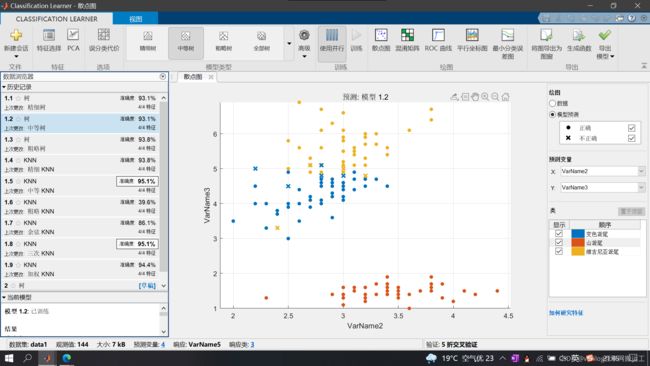
打开matlab机器学习工具箱选择决策树模型进行训练
选择准确度较高的决策树进行导出,点击右上角生成函数
导出后将已知变量进行替换(可以参考我ROC那篇博文)
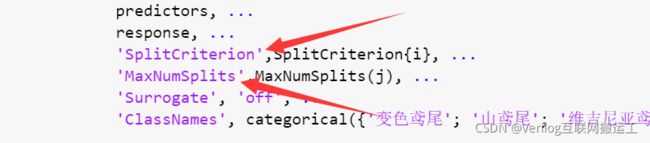
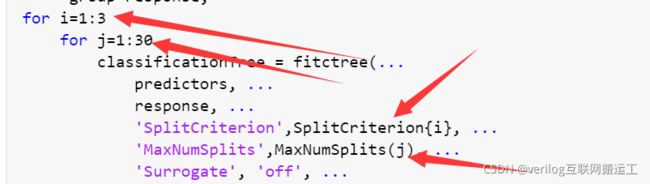
确定要调整的参数,本文以下图箭头所指两参数为例:
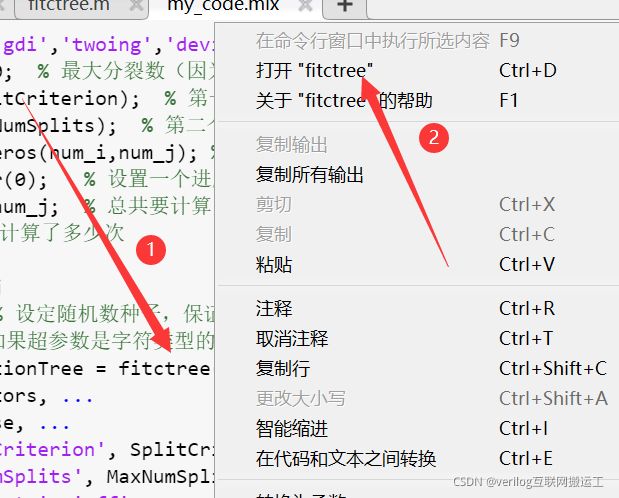
这两个参数分别为最大分裂数和分裂准则,为了找到具体的分裂准则,我们打开fitctree函数的内部,步骤如下:
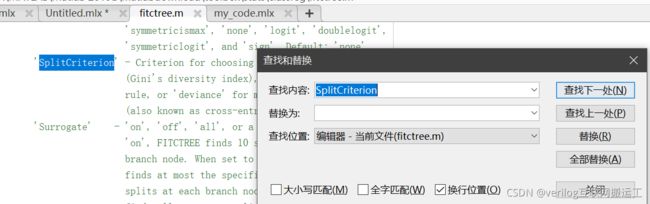
ctrl+F进行搜索即可:
然后将内容放入元胞数组中,并设置最大分裂次数的搜索范围为1到30:
在代码外层添加两个循环体i,j,然后修改参数:
修改完毕后再循环末尾储存每次的结果,红框内第一行为求混淆矩阵的函数,第二行用于求F1分数,其中F1分数位于stats结构体中microAVG的最后一行:
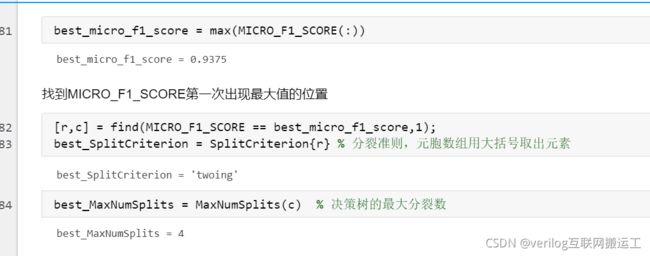
运行程序后寻找最大的F1分数的位置:
如果找的较好的F1分数,可在代码最前边加上rng随机数种子保存起来,否则下次运行会丢失结果。从上图中可以看出最大分裂次数为4、分裂准则为twoing时F1分数最好。
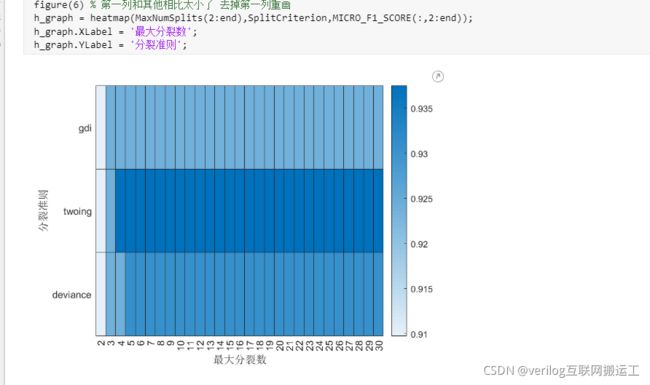
利用heatmap进行搜索可视化:
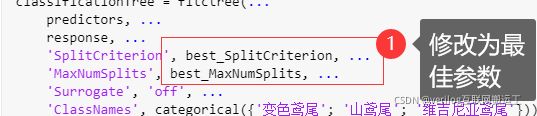
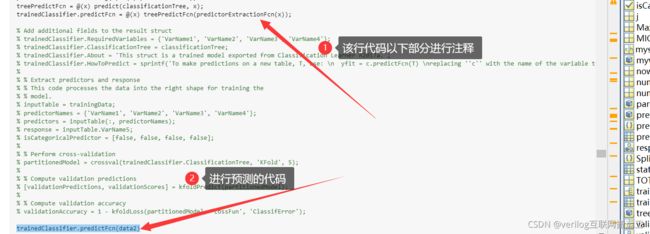
利用之前导出的函数在修改参数为最佳参数后进行预测,预测时将下图注释部分进行注释,然后加一行代码:trainedClassifier.predictFcn(data2) % data2为需要预测的数据
源代码(其中data1为训练集,data2为需要预测的数据):
%% 网格法过程
trainingData = data1;
inputTable = trainingData;
predictorNames = {'VarName1', 'VarName2', 'VarName3', 'VarName4'};
predictors = inputTable(:, predictorNames);
response = inputTable.VarName5;
isCategoricalPredictor = [false, false, false, false];
% 下面这两个变量在计算F1分数时会用到
ClassNames = unique(response); % y中各类的名称
group = response; % y所在的那一列(真实的类别)
% Train a classifier
% This code specifies all the classifier options and trains the classifier.
SplitCriterion = {'gdi','twoing','deviance'}; %分裂准则有三个可以选的,这里用元胞数组保存
MaxNumSplits = 1:30; % 最大分裂数(因为这个问题比较简单,我这里搜索设置的最大分裂数的上界是30)
num_i = length(SplitCriterion); % 第一个超参数SplitCriterion的可能性有3种
num_j = length(MaxNumSplits); % 第二个超参数MaxNumSplits的可能性有30种
MICRO_F1_SCORE = zeros(num_i,num_j); % 初始化最后得到的结果(初始化是为了加快代码运行速度)
mywaitbar = waitbar(0); % 设置一个进度条
TOTAL_NUM = num_i*num_j; % 总共要计算多少次
now_num = 0; % 已经计算了多少次
for i = 1:num_i
for j = 1:num_j
rng(520) % 设定随机数种子,保证结果的可重复性(这里的520也可以换成其他的数字)
% 注意,如果超参数是字符类型的话,它是被保存在元胞数组中的,取出时需要使用大括号才能得到字符型
classificationTree = fitctree(...
predictors, ...
response, ...
'SplitCriterion', SplitCriterion{i}, ...
'MaxNumSplits', MaxNumSplits(j), ...
'Surrogate', 'off', ...
'ClassNames', categorical({'变色鸢尾'; '山鸢尾'; '维吉尼亚鸢尾'}));
% 'SplitCriterion' - Criterion for choosing a split. One of 'gdi'
% (Gini's diversity index), 'twoing' for the twoing
% rule, or 'deviance' for maximum deviance reduction
% (also known as cross-entropy). Default: 'gdi'
% 'MaxNumSplits' - Maximal number of decision splits (or branch
% nodes) per tree. Default: size(X,1)-1
% Create the result struct with predict function
predictorExtractionFcn = @(t) t(:, predictorNames);
treePredictFcn = @(x) predict(classificationTree, x);
trainedClassifier.predictFcn = @(x) treePredictFcn(predictorExtractionFcn(x));
% Add additional fields to the result struct
trainedClassifier.RequiredVariables = {'VarName1', 'VarName2', 'VarName3', 'VarName4'};
trainedClassifier.ClassificationTree = classificationTree;
trainedClassifier.About = 'This struct is a trained model exported from Classification Learner R2017a.';
trainedClassifier.HowToPredict = sprintf('To make predictions on a new table, T, use: \n yfit = c.predictFcn(T) \nreplacing ''c'' with the name of the variable that is this struct, e.g. ''trainedModel''. \n \nThe table, T, must contain the variables returned by: \n c.RequiredVariables \nVariable formats (e.g. matrix/vector, datatype) must match the original training data. \nAdditional variables are ignored. \n \nFor more information, see How to predict using an exported model.');
% Extract predictors and response
% This code processes the data into the right shape for training the
% model.
inputTable = trainingData;
predictorNames = {'VarName1', 'VarName2', 'VarName3', 'VarName4'};
predictors = inputTable(:, predictorNames);
response = inputTable.VarName5;
isCategoricalPredictor = [false, false, false, false];
% Perform cross-validation
partitionedModel = crossval(trainedClassifier.ClassificationTree, 'KFold', 5);
% Compute validation predictions
[validationPredictions, validationScores] = kfoldPredict(partitionedModel);
% Compute validation accuracy
validationAccuracy = 1 - kfoldLoss(partitionedModel, 'LossFun', 'ClassifError');
% 下面就是计算每一次循环得到的micro F1分数
C = confusionmat(group,validationPredictions,'Order',ClassNames);
stats = statsOfMeasure(C);
MICRO_F1_SCORE(i,j) = stats.microAVG(end);
% 更新进度条
now_num = now_num+1;
mystr=['计算中...',num2str(100*now_num/TOTAL_NUM),'%'];
waitbar(now_num/TOTAL_NUM,mywaitbar,mystr);
end
end
%% 热力图
MICRO_F1_SCORE
% figure(5) % 画一个热力图
% heatmap(MaxNumSplits,SplitCriterion,MICRO_F1_SCORE)
figure(6) % 第一列和其他相比太小了 去掉第一列重画
h_graph = heatmap(MaxNumSplits(2:end),SplitCriterion,MICRO_F1_SCORE(:,2:end));
h_graph.XLabel = '最大分裂数';
h_graph.YLabel = '分裂准则';
%% 保存最佳位置
best_micro_f1_score = max(MICRO_F1_SCORE(:))
%找到MICRO_F1_SCORE第一次出现最大值的位置
[r,c] = find(MICRO_F1_SCORE == best_micro_f1_score,1);
best_SplitCriterion = SplitCriterion{r} % 分裂准则,元胞数组用大括号取出元素
best_MaxNumSplits = MaxNumSplits(c) % 决策树的最大分裂数
%%下面我们使用最优模型进行预测:
trainingData = data1;
inputTable = trainingData;
predictorNames = {'VarName1', 'VarName2', 'VarName3', 'VarName4'};
predictors = inputTable(:, predictorNames);
response = inputTable.VarName5;
isCategoricalPredictor = [false, false, false, false];
% Train a classifier
% This code specifies all the classifier options and trains the classifier.
classificationTree = fitctree(...
predictors, ...
response, ...
'SplitCriterion', best_SplitCriterion, ...
'MaxNumSplits', best_MaxNumSplits, ...
'Surrogate', 'off', ...
'ClassNames', categorical({'变色鸢尾'; '山鸢尾'; '维吉尼亚鸢尾'}));
% Create the result struct with predict function
predictorExtractionFcn = @(t) t(:, predictorNames);
treePredictFcn = @(x) predict(classificationTree, x);
trainedClassifier.predictFcn = @(x) treePredictFcn(predictorExtractionFcn(x));
% Add additional fields to the result struct
% trainedClassifier.RequiredVariables = {'VarName1', 'VarName2', 'VarName3', 'VarName4'};
% trainedClassifier.ClassificationTree = classificationTree;
% trainedClassifier.About = 'This struct is a trained model exported from Classification Learner R2017a.';
% trainedClassifier.HowToPredict = sprintf('To make predictions on a new table, T, use: \n yfit = c.predictFcn(T) \nreplacing ''c'' with the name of the variable that is this struct, e.g. ''trainedModel''. \n \nThe table, T, must contain the variables returned by: \n c.RequiredVariables \nVariable formats (e.g. matrix/vector, datatype) must match the original training data. \nAdditional variables are ignored. \n \nFor more information, see How to predict using an exported model.');
%
% % Extract predictors and response
% % This code processes the data into the right shape for training the
% % model.
% inputTable = trainingData;
% predictorNames = {'VarName1', 'VarName2', 'VarName3', 'VarName4'};
% predictors = inputTable(:, predictorNames);
% response = inputTable.VarName5;
% isCategoricalPredictor = [false, false, false, false];
%
% % Perform cross-validation
% partitionedModel = crossval(trainedClassifier.ClassificationTree, 'KFold', 5);
%
% % Compute validation predictions
% [validationPredictions, validationScores] = kfoldPredict(partitionedModel);
%
% % Compute validation accuracy
% validationAccuracy = 1 - kfoldLoss(partitionedModel, 'LossFun', 'ClassifError');
predict_final_result=trainedClassifier.predictFcn(data2);%预测的结果参考:代码参考数学建模清风老师的代码,本文章仅用于整理分享学习笔记,无任何商业用途,若有意见可联系本人删除