投稿在其他公众号里了,这份为作死测试用的,真的不是抄袭,,,
karyoploteR
- official website(github)
- bioconductor
- github
- 帮助文档HTML版
- pubmed文章
简介
一个用来做多物种染色体分布情况图的R包,可以非常方便的添加多种自定义内容,使用体验,爽
一、 基础用法
1. 最基础的图形
kp <- plotKaryotype(genome="mm10", plot.type=1, main="The mm10 genome", cex=0.6)
- genome:指定要做什么染色体组的图,eg:mm10、hg19
- plot.type:有1,2,3,4,5五种,在染色体形状的细节上有区别,可以根据自己的喜好修改
- main:标题
- cex:字体比例,cex越大字体越大
效果如下:
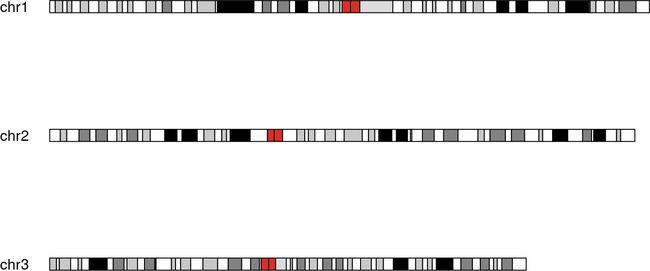
2. 只做某几个染色体的
kp <- plotKaryotype(genome="hg19", plot.type=2, chromosomes=c("chr1", "chr2", "chr3"))
效果如下:红色标记为plot.type 2和1的主要区别
3. 添加附加的坐标轴
kp <- plotKaryotype(chromosomes=c("chr1", "chr2"), plot.type=2)
# data.panel=1,添加在染色体上方
kpDataBackground(kp)
# data.panel=2,添加在染色体下方,添加了两个data.panel
kpDataBackground(kp, r1=0.47, data.panel=2)
kpDataBackground(kp, r0=0.53, data.panel=2)
# 开始添加坐标轴
# Default axis,添加染色体上方的左侧轴
kpAxis(kp)
# Axis on the right side of the data.panel,添加染色体上方的右侧轴
kpAxis(kp, side = 2)
# Changing the limits and having more ticks, with a smaller font size,调整染色体下方第一个panel的左侧轴
kpAxis(kp, r1=0.47, ymin=-5000, ymax = 5000, numticks = 5, cex=0.5, data.panel=2)
#and a different scale on the right,调整染色体下方第一个panel的右侧轴
kpAxis(kp, r1=0.47, ymin=-2, ymax = 2, numticks = 3, cex=0.5, data.panel=2, side=2)
#Changing the colors and labels and tick positions,调整染色体下方第二个panel的左侧轴
kpAxis(kp, r0=0.53, tick.pos = c(0.3, 0.6, 1), labels = c("A", "B", "C"), col="#66AADD",
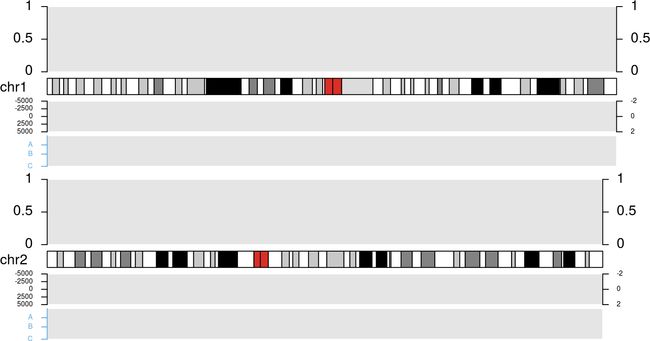
cex=0.5, data.panel=2)
效果如下:上方代码是官方文档里的,只不过我调整了顺序,可能更便于理解
4. 添加附加的散点图
# 生成散点图需要的点
data.points <- data.frame(chr="chr1", pos=(1:240)*1e6, value=rnorm(240, 0.5, 0.1))
# 只做一个染色体的内容
kp <- plotKaryotype(plot.type = 4, chromosomes = "chr1")
# 添加一个附加图的背景,data.panel为1,应当是在染色体上方作图的意思
kpDataBackground(kp, data.panel = 1)
# 添加染色体上的距离标记
kpAddBaseNumbers(kp)
# 做散点图
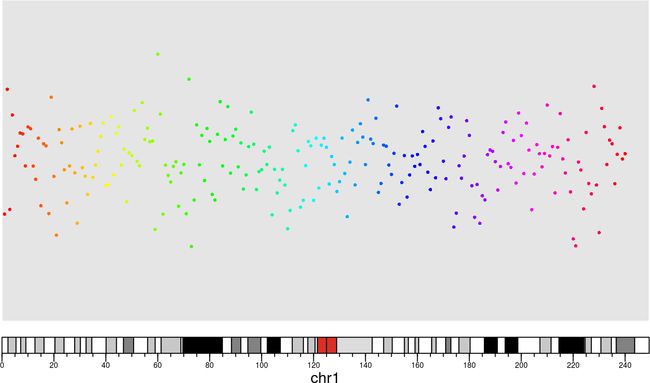
kpPoints(kp, chr = data.points$chr, x=data.points$pos, y=data.points$value, col=rainbow(240))
效果如下:
5. 添加自定义的区段
这个也是用的最多的一个了吧,相当简单,从DataFrame直接往里边添加即可
dataframe只需要包含三列内容:chr、start、end
# 第一个自定义的区段
gains <- makeGRangesFromDataFrame(data.frame(chr=c("chr1", "chr5", "chr17", "chr22"), start=c(1, 1000000, 4000000, 1), end=c(5000000, 3200000, 80000000, 1200000)))
# 第二个自顶一个区段
losses <- makeGRangesFromDataFrame(data.frame(chr=c("chr3", "chr9", "chr17"), start=c(80000000, 20000000, 1), end=c(170000000, 30000000, 25000000)))
# 作图,并且以特定的颜色添加自定义的区段
kp <- plotKaryotype(genome="hg19")
kpPlotRegions(kp, gains, col="#FFAACC")
kpPlotRegions(kp, losses, col="#CCFFAA")
效果如下:
如果自定义的染色体位置中间包含有重合位点,会自动在同一个位置,用几个不同的段来展示,如下:
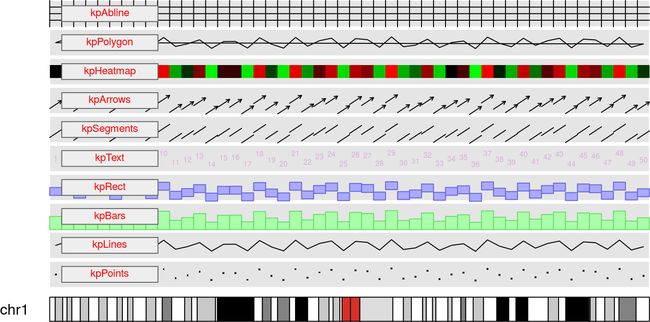
6. 最后,扔一段集大成的代码和图,供大家参考
pp <- getDefaultPlotParams(plot.type = 1)
pp$data1height=600
tr.i <- 1/11
tr.o <- 1/10
kp <- plotKaryotype(chromosomes=c("chr1"), plot.params = pp)
dd <- toGRanges(data.frame(chr="chr1", start=end(kp$genome[1])/50*(0:49), end=end(kp$genome[1])/50*(1:50)))
mcols(dd) <- data.frame(y=((sin(start(dd)) + rnorm(n=50, mean=0, sd=0.1))/5)+0.5)
tn <- 0
kpDataBackground(kp, r0=tr.o*tn, r1=tr.o*tn+tr.i)
kpPoints(kp, dd, r0=tr.o*tn, r1=tr.o*tn+tr.i, pch=".", cex=2)
kpRect(kp, chr="chr1", x0=5000000, x1=45000000, y0=0.2, y1=0.8, r0=tr.o*tn, r1=tr.o*tn+tr.i, col="#EEEEEE", border="#666666")
kpText(kp, chr="chr1", x=25000000, y=0.5, col="red", r0=tr.o*tn, r1=tr.o*tn+tr.i, labels="kpPoints", cex=0.7)
tn <- 1
kpDataBackground(kp, r0=tr.o*tn, r1=tr.o*tn+tr.i)
kpLines(kp, dd, r0=tr.o*tn, r1=tr.o*tn+tr.i, pch=".", cex=2)
kpRect(kp, chr="chr1", x0=5000000, x1=45000000, y0=0.2, y1=0.8, r0=tr.o*tn, r1=tr.o*tn+tr.i, col="#EEEEEE", border="#666666")
kpText(kp, chr="chr1", x=25000000, y=0.5, col="red", r0=tr.o*tn, r1=tr.o*tn+tr.i, labels="kpLines", cex=0.7)
tn <- 2
kpDataBackground(kp, r0=tr.o*tn, r1=tr.o*tn+tr.i)
kpBars(kp, dd, y1=dd$y, r0=tr.o*tn, r1=tr.o*tn+tr.i, col="#AAFFAA", border="#66DD66")
kpRect(kp, chr="chr1", x0=5000000, x1=45000000, y0=0.2, y1=0.8, r0=tr.o*tn, r1=tr.o*tn+tr.i, col="#EEEEEE", border="#666666")
kpText(kp, chr="chr1", x=25000000, y=0.5, col="red", r0=tr.o*tn, r1=tr.o*tn+tr.i, labels="kpBars", cex=0.7)
tn <- 3
kpDataBackground(kp, r0=tr.o*tn, r1=tr.o*tn+tr.i)
kpRect(kp, dd, y0=dd$y-0.3, y1=dd$y, r0=tr.o*tn, r1=tr.o*tn+tr.i, col="#AAAAFF", border="#6666DD")
kpRect(kp, chr="chr1", x0=5000000, x1=45000000, y0=0.2, y1=0.8, r0=tr.o*tn, r1=tr.o*tn+tr.i, col="#EEEEEE", border="#666666")
kpText(kp, chr="chr1", x=25000000, y=0.5, col="red", r0=tr.o*tn, r1=tr.o*tn+tr.i, labels="kpRect", cex=0.7)
tn <- 4
kpDataBackground(kp, r0=tr.o*tn, r1=tr.o*tn+tr.i)
kpText(kp, dd, labels=as.character(1:50), cex=0.5, r0=tr.o*tn, r1=tr.o*tn+tr.i, col="#DDAADD")
kpRect(kp, chr="chr1", x0=5000000, x1=45000000, y0=0.2, y1=0.8, r0=tr.o*tn, r1=tr.o*tn+tr.i, col="#EEEEEE", border="#666666")
kpText(kp, chr="chr1", x=25000000, y=0.5, col="red", r0=tr.o*tn, r1=tr.o*tn+tr.i, labels="kpText", cex=0.7)
tn <- 5
kpDataBackground(kp, r0=tr.o*tn, r1=tr.o*tn+tr.i)
kpSegments(kp, dd, y0=dd$y-0.3, y1=dd$y, r0=tr.o*tn, r1=tr.o*tn+tr.i)
kpRect(kp, chr="chr1", x0=5000000, x1=45000000, y0=0.2, y1=0.8, r0=tr.o*tn, r1=tr.o*tn+tr.i, col="#EEEEEE", border="#666666")
kpText(kp, chr="chr1", x=25000000, y=0.5, col="red", r0=tr.o*tn, r1=tr.o*tn+tr.i, labels="kpSegments", cex=0.7)
tn <- 6
kpDataBackground(kp, r0=tr.o*tn, r1=tr.o*tn+tr.i)
kpArrows(kp, dd, y0=dd$y-0.3, y1=dd$y, r0=tr.o*tn, r1=tr.o*tn+tr.i, length=0.04)
kpRect(kp, chr="chr1", x0=5000000, x1=45000000, y0=0.2, y1=0.8, r0=tr.o*tn, r1=tr.o*tn+tr.i, col="#EEEEEE", border="#666666")
kpText(kp, chr="chr1", x=25000000, y=0.5, col="red", r0=tr.o*tn, r1=tr.o*tn+tr.i, labels="kpArrows", cex=0.7)
tn <- 7
kpDataBackground(kp, r0=tr.o*tn, r1=tr.o*tn+tr.i)
kpHeatmap(kp, dd, r0=tr.o*tn+tr.i/4, r1=tr.o*tn+tr.i-tr.i/4, colors = c("green", "black", "red"))
kpRect(kp, chr="chr1", x0=5000000, x1=45000000, y0=0.2, y1=0.8, r0=tr.o*tn, r1=tr.o*tn+tr.i, col="#EEEEEE", border="#666666")
kpText(kp, chr="chr1", x=25000000, y=0.5, col="red", r0=tr.o*tn, r1=tr.o*tn+tr.i, labels="kpHeatmap", cex=0.7)
tn <- 8
kpDataBackground(kp, r0=tr.o*tn, r1=tr.o*tn+tr.i)
kpPolygon(kp, dd, r0=tr.o*tn, r1=tr.o*tn+tr.i)
kpRect(kp, chr="chr1", x0=5000000, x1=45000000, y0=0.2, y1=0.8, r0=tr.o*tn, r1=tr.o*tn+tr.i, col="#EEEEEE", border="#666666")
kpText(kp, chr="chr1", x=25000000, y=0.5, col="red", r0=tr.o*tn, r1=tr.o*tn+tr.i, labels="kpPolygon", cex=0.7)
tn <- 9
kpDataBackground(kp, r0=tr.o*tn, r1=tr.o*tn+tr.i)
kpAbline(kp, h=c(0.25, 0.5, 0.75), v=start(dd), r0=tr.o*tn, r1=tr.o*tn+tr.i)
kpRect(kp, chr="chr1", x0=5000000, x1=45000000, y0=0.2, y1=0.8, r0=tr.o*tn, r1=tr.o*tn+tr.i, col="#EEEEEE", border="#666666")
kpText(kp, chr="chr1", x=25000000, y=0.5, col="red", r0=tr.o*tn, r1=tr.o*tn+tr.i, labels="kpAbline", cex=0.7)