Python:S2实现
title={S2: An efficient graph based active learning algorithm with application to nonparametric classification},
author={Dasarathy, Gautam and Nowak, Robert and Zhu, Xiaojin}
import networkx as nx
import matplotlib.pyplot as plt
from venv.S2 import s2, path_midpoint, enumerate_find_ssp
from venv.S2.moss import moss
# from venv.S2.util import draw_labeled_graph
import timeit
from sklearn import datasets
from scipy.spatial.distance import pdist, squareform
import numpy as np
from copy import deepcopy
def draw_labeled_graph(G, oracle):
def label_to_color(l):
if l is None: return '0.75'
return 'r' if l > 0 else 'b'
nx.draw(G,
pos={n: n for n in G.nodes()},
node_color=[label_to_color(oracle(n)) for n in G.nodes()])
def test_simple_lattice():
# G = nx.grid_2d_graph(10, 10)
# X, y = datasets.load_iris(return_X_y=True)
X, y = datasets.make_blobs(n_samples=200, n_features=2, centers=2, cluster_std=[3, 3], random_state=1)
N = X.shape[0]
distlist = pdist(X, metric='euclidean')
dist_Matrix = squareform(distlist)
simi_Matrix = np.zeros((N, N))
neiNum = 5
G = nx.Graph()
for i in range(N):
ordidx = np.argsort(dist_Matrix[i, :])
for j in range(neiNum + 1):
if i != ordidx[j]:
simi_Matrix[i, ordidx[j]] = dist_Matrix[i, ordidx[j]]
for i in range(N):
for j in range(N):
if simi_Matrix[i, j] > 0:
G.add_weighted_edges_from([(i, j, simi_Matrix[i, j])])
def oracle(vert):
if y[vert] == 1:
return True
else:
return False
# def oracle(vert):
# return ((vert[0] < 3) and (vert[1] < 3)) or ((vert[0] > 6) and (vert[1] > 6))
# enum: 638 ms ± 22.8 ms per loop (mean ± std. dev. of 7 runs, 1 loop each)
# moss: 18.1 ms ± 75.7 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)
G_cut = s2(G, oracle, lambda G, U, V: moss(G, U, V))
fig = plt.figure()
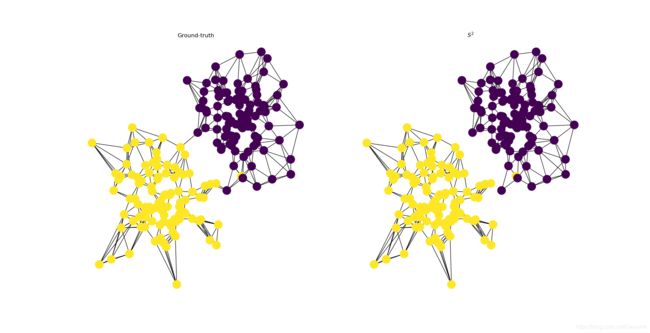
fig.add_subplot(121).title.set_text('Ground-truth')
nodes = np.array([_ for _ in range(N)])
vnode = deepcopy(X)
npos = dict(zip(nodes,vnode))
pos = {}
pos.update(npos)
nx.draw(G,pos,node_color=y)
# draw_labeled_graph(G, oracle)
fig.add_subplot(122).title.set_text('$S^2$')
nx.draw(G_cut,pos, node_color=y)
# draw_labeled_graph(G_cut, lambda v: G_cut.node[v].get('label'))
plt.show()
if __name__ == '__main__':
test_simple_lattice()
from collections import deque
import networkx as nx
def moss(G, U, V):
queue_u, queue_v = deque([]), deque([])
visited_u, visited_v = set(), set()
for u in U:
queue_u.append((u, G.neighbors(u)))
visited_u.add(u)
for v in V:
queue_v.append((v, G.neighbors(v)))
visited_v.add(v)
while queue_u and queue_v:
parent, children = queue_u.popleft()
for child in children:
if child not in visited_u:
visited_u.add(child)
queue_u.append((child, G.neighbors(child)))
if child in visited_v and child not in V:
return child
parent, children = queue_v.popleft()
for child in children:
if child not in visited_v:
visited_v.add(child)
queue_v.append((child, G.neighbors(child)))
if child in visited_u and child not in U:
return child
参考:https://github.com/erinzm/s2/blob/master/s2/__init__.py