三维点云学习(9)5-实现RANSAC Registration配准
三维点云学习(9)5-实现RANSAC Registration配准
参考博客:
机器视觉之 ICP算法和RANSAC算法
三维点云配准
ICP点云配准原理及优化
本章因个人能力有限,大部分代码摘自github大神的code
数据集
链接:https://pan.baidu.com/s/11OhSCA2Ck-WA6_b-iCNsfA
提取码:hyi3
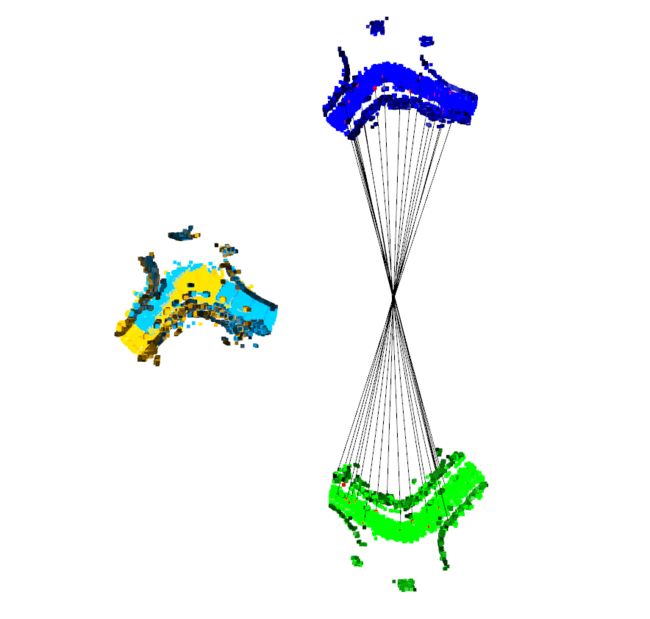
效果图:
本次以 数据集 643.bin ; 456.bin为例
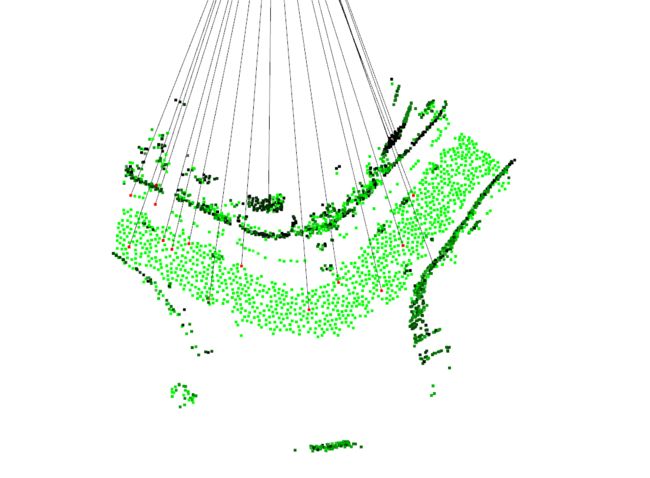
蓝色和绿色分别为 source 点云图和 target 点云图;彩色为经过配准后拼接效果图,黑色为 特征点与特征点之间的连线




189.bin 270.bin


编码思路:
step1 读取数据
step2 对点云进行降采样downsample
step3 iss 特征点提取
step4 fpfh特征点描述 feature description
step5 RANSAC Registration,初配准、得到初始旋转、平移矩阵
step5.1 Establish correspondences(point pairs) 建立 pairs
step5.2 select 4 pairs at each iteration,选择4对corresponding 进行模型拟合
step5.3 iter 迭代, iter_match() ,选择出 vaild T
step6 ICP for refined estimation,优化配准 ICP对初始T进行迭代优化:
部分编码模块
svd 求解选准平移矩阵
def solve_procrustes_transf(P,Q): # 求解平移旋转矩阵 solve_procrustes_transformation
up = P.mean(axis=0)
uq = Q.mean(axis=0)
# move to center:
P_centered = P - up
Q_centered = Q - uq
U, s, V = np.linalg.svd(np.dot(Q_centered.T, P_centered), full_matrices=True, compute_uv=True)
R = np.dot(U, V)
t = uq - np.dot(R, up)
# format as transform:
T = np.zeros((4, 4))
T[0:3, 0:3] = R
T[0:3, 3] = t
T[3, 3] = 1.0
return T
ransac 初配准
def ransac_match(
idx_target,idx_source,
pcd_source, pcd_target,
feature_source, feature_target,
ransac_params, checker_params
):
#step5.1 Establish correspondences(point pairs) 建立 pairs
matches = get_init_matches(feature_source, feature_target) #通过 fpfh 建立的feature squre map 建立最初的 pairs
##build search tree on the target:
search_tree_target = o3d.geometry.KDTreeFlann(pcd_target)
N, _ = matches.shape
idx_matches = np.arange(N) #对每队 pair 打标签
T = None #translation martix
#step5.2 select 4 pairs at each iteration,选择4对corresponding 进行模型拟合
proposal_generator = (
matches[np.random.choice(idx_matches, ransac_params.num_samples, replace=False)] for _ in iter(int, 1)
)
##step5.3 iter 迭代, iter_match() ,选择出 vaild T
validator = lambda proposal: iter_match(idx_target,idx_source,pcd_source, pcd_target, proposal, checker_params)
with concurrent.futures.ThreadPoolExecutor(max_workers=ransac_params.max_workers) as executor:
for T in map(
validator,
proposal_generator #map()是 Python 内置的高阶函数,它接收一个函数 f 和一个 list,并通过把函数 f 依次作用在 list 的每个元素上,得到一个新的 list 并返回。
):
print(T)
if not (T is None):
break
#set baseline
print('[RANSAC ICP]: Get first valid proposal. Start registration...')
best_result = icp_exact_match(
pcd_source, pcd_target, search_tree_target,
T,
ransac_params.max_correspondence_distance,
ransac_params.max_refinement
)
# RANSAC:
num_validation = 0
for i in range(ransac_params.max_iteration):
# get proposal:
T = validator(next(proposal_generator))
# check validity:
if (not (T is None)) and (num_validation < ransac_params.max_validation):
num_validation += 1
# refine estimation on all keypoints:
result = icp_exact_match(
pcd_source, pcd_target, search_tree_target,
T,
ransac_params.max_correspondence_distance,
ransac_params.max_refinement
)
# update best result:
best_result = best_result if best_result.fitness > result.fitness else result
if num_validation == ransac_params.max_validation:
break
return best_result
ICP配准优化
def icp_exact_match(
pcd_source, pcd_target, search_tree_target,
T,
max_correspondence_distance, max_iteration
):
# num. points in the source:
N = len(pcd_source.points)
# evaluate relative change for early stopping:
result_prev = result_curr = o3d.registration.evaluate_registration(
pcd_source, pcd_target, max_correspondence_distance, T
)
for _ in range(max_iteration):
# TODO: transform is actually an in-place operation. deep copy first otherwise the result will be WRONG
pcd_source_current = copy.deepcopy(pcd_source)
# apply transform:
pcd_source_current = pcd_source_current.transform(T)
# find correspondence:
matches = []
for n in range(N):
query = np.asarray(pcd_source_current.points)[n]
_, idx_nn_target, dis_nn_target = search_tree_target.search_knn_vector_3d(query, 1)
if dis_nn_target[0] <= max_correspondence_distance:
matches.append(
[n, idx_nn_target[0]]
)
matches = np.asarray(matches)
#icp
if len(matches) >= 4:
# sovle ICP:
P = np.asarray(pcd_source.points)[matches[:, 0]]
Q = np.asarray(pcd_target.points)[matches[:, 1]]
T = solve_procrustes_transf(P, Q)
# evaluate:
result_curr = o3d.registration.evaluate_registration(
pcd_source, pcd_target, max_correspondence_distance, T
)
# if no significant improvement:提前中止
if shall_terminate(result_curr, result_prev):
print('[RANSAC ICP]: Early stopping.')
break
return result_curr
全部代码:(utils/io.py、utils/visualize.py、main.py、RANSAC.py、ICP.pyiss.py)
import os
import argparse
import progressbar
import collections
import numpy as np
import open3d as o3d
#IO utils:
import utils.io as io
import utils.visualize as visualize
import matplotlib.pyplot as plt
#导入iss feature detection 模块
from iss import detect
from RANSAC import ransac_match
from ICP import icp_exact_match
#RANSAC configuration:
RANSACParams = collections.namedtuple(
'RANSACParams',
[
'max_workers',
'num_samples',
'max_correspondence_distance', 'max_iteration', 'max_validation', 'max_refinement'
]
)
# fast pruning algorithm configuration:
CheckerParams = collections.namedtuple(
'CheckerParams',
['max_correspondence_distance', 'max_edge_length_ratio', 'normal_angle_threshold']
)
if __name__ == '__main__':
radius = 0.5
#step1 读取数据
input_dir = "registration_dataset"
registration_results = io.read_registration_results(os.path.join(input_dir,'reg_result.txt')) #读取 reg_result.txt 结果
##init output
df_output = io.init_output()
##批量读取
# for i,r in (
# list(registration_results.iterrows())
# ):
# idx_target = int(r['idx1'])
# idx_source = int(r['idx2'])
idx_target = 643
idx_source = 456
##load point cloud
pcd_source = io.read_point_cloud_bin(
os.path.join(input_dir,'point_clouds',f'{idx_source}.bin')
)
pcd_target = io.read_point_cloud_bin(
os.path.join(input_dir,'point_clouds',f'{idx_target}.bin')
)
#step2 对点云进行降采样downsample
pcd_source,idx_inliers = pcd_source.remove_radius_outlier(nb_points=4,radius=radius)
pcd_target, idx_inliers = pcd_target.remove_radius_outlier(nb_points=4, radius=radius)
## 构建 kd-tree
search_tree_source = o3d.geometry.KDTreeFlann(pcd_source)
search_tree_target = o3d.geometry.KDTreeFlann(pcd_target)
#step3 iss 特征点提取
keypoints_source = detect(pcd_source, search_tree_source, radius)
keypoints_target = detect(pcd_target, search_tree_target, radius)
#step4 fpfh特征点描述 feature description
pcd_source_keypoints = pcd_source.select_by_index(keypoints_source['id'].values)
##fpfh 进行 特征点描述
fpfh_source_keypoints = o3d.registration.compute_fpfh_feature(
pcd_source_keypoints,
o3d.geometry.KDTreeSearchParamHybrid(radius=radius*5,max_nn=100)
).data
pcd_target_keypoints = pcd_target.select_by_index(keypoints_target['id'].values)
fpfh_target_keypoints = o3d.registration.compute_fpfh_feature(
pcd_target_keypoints,
o3d.geometry.KDTreeSearchParamHybrid(radius=radius*5,max_nn=100)
).data
# generate matches:
distance_threshold_init = 1.5 * radius
distance_threshold_final = 1.0 * radius
#step5 RANSAC Registration,初配准、得到初始旋转、平移矩阵
init_result = ransac_match(
idx_target,idx_source,
pcd_source_keypoints, pcd_target_keypoints,
fpfh_source_keypoints, fpfh_target_keypoints,
ransac_params=RANSACParams(
max_workers=5,
num_samples=4,
max_correspondence_distance=distance_threshold_init,
max_iteration=200000,
max_validation=500,
max_refinement=30
),
checker_params=CheckerParams(
max_correspondence_distance=distance_threshold_init,
max_edge_length_ratio=0.9,
normal_angle_threshold=None
)
)
#step6 ICP for refined estimation,优化配准 ICP对初始T进行迭代优化:
final_result = icp_exact_match(
pcd_source, pcd_target, search_tree_target,
init_result.transformation,
distance_threshold_final, 60
)
# visualize:
visualize.show_registration_result(
pcd_source_keypoints, pcd_target_keypoints, init_result.correspondence_set,
pcd_source, pcd_target, final_result.transformation
)
#
# # add result:
# io.add_to_output(df_output, idx_target, idx_source, final_result.transformation)
#
# # write output:
# io.write_output(
# os.path.join(input_dir, 'reg_result_yaogefad.txt'),
# df_output
# )
RANSAC.py
import os
import argparse
import progressbar
import collections
import numpy as np
import open3d as o3d
import copy
import concurrent.futures
from scipy.spatial.distance import pdist
from scipy.spatial.transform import Rotation
#IO utils:
import utils.io as io
import utils.visualize as visualize
import matplotlib.pyplot as plt
from ICP import icp_exact_match
def solve_procrustes_transf(P,Q): # 求解平移旋转矩阵 solve_procrustes_transformation
up = P.mean(axis=0)
uq = Q.mean(axis=0)
# move to center:
P_centered = P - up
Q_centered = Q - uq
U, s, V = np.linalg.svd(np.dot(Q_centered.T, P_centered), full_matrices=True, compute_uv=True)
R = np.dot(U, V)
t = uq - np.dot(R, up)
# format as transform:
T = np.zeros((4, 4))
T[0:3, 0:3] = R
T[0:3, 3] = t
T[3, 3] = 1.0
return T
def get_init_matches(feature_source, feature_target):
##对 feature target fpfh 建立 kd—tree
fpfh_search_tree = o3d.geometry.KDTreeFlann(feature_target)
##建立 pairs
_,N = feature_source.shape
matches = []
for i in range(N):
query = feature_source[:,i]
_, idx_nn_target, _ = fpfh_search_tree.search_knn_vector_xd(query, 1) #source -> target
matches.append([i,idx_nn_target[0]]) #通过knn 寻找唯一 的 nearest points 一一配对 构建pair
matches = np.asarray(matches)
return matches
def iter_match(
idx_target,idx_source,
pcd_source, pcd_target,
proposal,
checker_params
):
idx_source, idx_target = proposal[:, 0], proposal[:, 1]
#法向量校准
if not checker_params.normal_angle_threshold is None:
# get corresponding normals:
normals_source = np.asarray(pcd_source.normals)[idx_source]
normals_target = np.asarray(pcd_target.normals)[idx_target]
# a. normal direction check:
normal_cos_distances = (normals_source*normals_target).sum(axis = 1)
is_valid_normal_match = np.all(normal_cos_distances >= np.cos(checker_params.normal_angle_threshold))
if not is_valid_normal_match:
return None
# get corresponding points:
points_source = np.asarray(pcd_source.points)[idx_source]
points_target = np.asarray(pcd_target.points)[idx_target]
# b. edge length ratio check:
#构建距离矩阵,使用 Mutual nearest descriptor matching
pdist_source = pdist(points_source)
pdist_target = pdist(points_target)
is_valid_edge_length = np.all(
np.logical_and(
pdist_source > checker_params.max_edge_length_ratio * pdist_target,
pdist_target > checker_params.max_edge_length_ratio * pdist_source
)
)
if not is_valid_edge_length:
return None
# c. fast correspondence distance check:s
T = solve_procrustes_transf(points_source, points_target) #通过 svd 初步求解 旋转、平移矩阵
R, t = T[0:3, 0:3], T[0:3, 3]
#deviation:偏差 区分 inline outline 通过 距离判断
deviation = np.linalg.norm(
points_target - np.dot(points_source, R.T) - t,
axis = 1
)
#判断数目
is_valid_correspondence_distance = np.all(deviation <= checker_params.max_correspondence_distance)
return T if is_valid_correspondence_distance else None
def ransac_match(
idx_target,idx_source,
pcd_source, pcd_target,
feature_source, feature_target,
ransac_params, checker_params
):
#step5.1 Establish correspondences(point pairs) 建立 pairs
matches = get_init_matches(feature_source, feature_target) #通过 fpfh 建立的feature squre map 建立最初的 pairs
##build search tree on the target:
search_tree_target = o3d.geometry.KDTreeFlann(pcd_target)
N, _ = matches.shape
idx_matches = np.arange(N) #对每队 pair 打标签
T = None #translation martix
#step5.2 select 4 pairs at each iteration,选择4对corresponding 进行模型拟合
proposal_generator = (
matches[np.random.choice(idx_matches, ransac_params.num_samples, replace=False)] for _ in iter(int, 1)
)
##step5.3 iter 迭代, iter_match() ,选择出 vaild T
validator = lambda proposal: iter_match(idx_target,idx_source,pcd_source, pcd_target, proposal, checker_params)
with concurrent.futures.ThreadPoolExecutor(max_workers=ransac_params.max_workers) as executor:
for T in map(
validator,
proposal_generator #map()是 Python 内置的高阶函数,它接收一个函数 f 和一个 list,并通过把函数 f 依次作用在 list 的每个元素上,得到一个新的 list 并返回。
):
print(T)
if not (T is None):
break
#set baseline
print('[RANSAC ICP]: Get first valid proposal. Start registration...')
best_result = icp_exact_match(
pcd_source, pcd_target, search_tree_target,
T,
ransac_params.max_correspondence_distance,
ransac_params.max_refinement
)
# RANSAC:
num_validation = 0
for i in range(ransac_params.max_iteration):
# get proposal:
T = validator(next(proposal_generator))
# check validity:
if (not (T is None)) and (num_validation < ransac_params.max_validation):
num_validation += 1
# refine estimation on all keypoints:
result = icp_exact_match(
pcd_source, pcd_target, search_tree_target,
T,
ransac_params.max_correspondence_distance,
ransac_params.max_refinement
)
# update best result:
best_result = best_result if best_result.fitness > result.fitness else result
if num_validation == ransac_params.max_validation:
break
return best_result
ICP.py
import os
import argparse
import progressbar
import collections
import numpy as np
import open3d as o3d
import copy
import concurrent.futures
from scipy.spatial.distance import pdist
from scipy.spatial.transform import Rotation
#IO utils:
import utils.io as io
import utils.visualize as visualize
import matplotlib.pyplot as plt
def solve_procrustes_transf(P,Q): # 求解平移旋转矩阵 solve_procrustes_transformation
up = P.mean(axis=0)
uq = Q.mean(axis=0)
# move to center:
P_centered = P - up
Q_centered = Q - uq
U, s, V = np.linalg.svd(np.dot(Q_centered.T, P_centered), full_matrices=True, compute_uv=True)
R = np.dot(U, V)
t = uq - np.dot(R, up)
# format as transform:
T = np.zeros((4, 4))
T[0:3, 0:3] = R
T[0:3, 3] = t
T[3, 3] = 1.0
return T
def shall_terminate(result_curr, result_prev):
# relative fitness improvement:
relative_fitness_gain = result_curr.fitness / result_prev.fitness - 1
return relative_fitness_gain < 0.01
def icp_exact_match(
pcd_source, pcd_target, search_tree_target,
T,
max_correspondence_distance, max_iteration
):
# num. points in the source:
N = len(pcd_source.points)
# evaluate relative change for early stopping:
result_prev = result_curr = o3d.registration.evaluate_registration(
pcd_source, pcd_target, max_correspondence_distance, T
)
for _ in range(max_iteration):
# TODO: transform is actually an in-place operation. deep copy first otherwise the result will be WRONG
pcd_source_current = copy.deepcopy(pcd_source)
# apply transform:
pcd_source_current = pcd_source_current.transform(T)
# find correspondence:
matches = []
for n in range(N):
query = np.asarray(pcd_source_current.points)[n]
_, idx_nn_target, dis_nn_target = search_tree_target.search_knn_vector_3d(query, 1)
if dis_nn_target[0] <= max_correspondence_distance:
matches.append(
[n, idx_nn_target[0]]
)
matches = np.asarray(matches)
#icp
if len(matches) >= 4:
# sovle ICP:
P = np.asarray(pcd_source.points)[matches[:, 0]]
Q = np.asarray(pcd_target.points)[matches[:, 1]]
T = solve_procrustes_transf(P, Q)
# evaluate:
result_curr = o3d.registration.evaluate_registration(
pcd_source, pcd_target, max_correspondence_distance, T
)
# if no significant improvement:提前中止
if shall_terminate(result_curr, result_prev):
print('[RANSAC ICP]: Early stopping.')
break
return result_curr
io.py
import numpy as np
import pandas as pd
import open3d as o3d
from scipy.spatial.transform import Rotation as R
def read_point_cloud_bin(bin_path):
"""
Read point cloud in bin format
Parameters
----------
bin_path: str
Input path of Oxford point cloud bin
Returns
----------
"""
data = np.fromfile(bin_path, dtype=np.float32)
# format:
N, D = data.shape[0]// 6, 6
point_cloud_with_normal = np.reshape(data, (N, D))
point_cloud = o3d.geometry.PointCloud()
point_cloud.points = o3d.utility.Vector3dVector(point_cloud_with_normal[:, 0:3])
point_cloud.normals = o3d.utility.Vector3dVector(point_cloud_with_normal[:, 3:6])
return point_cloud
def read_registration_results(results_path):
"""
Read
Parameters
----------
results_path: str
Input path of point cloud registration results
Returns
----------
"""
# load csv:
df_results = pd.read_csv(
results_path
)
return df_results
def init_output():
"""
Get registration result output template
"""
df_output = {
'idx1': [],
'idx2': [],
't_x': [],
't_y': [],
't_z': [],
'q_w': [],
'q_x': [],
'q_y': [],
'q_z': []
}
return df_output
def add_to_output(df_output, idx1, idx2, T):
"""
Add record to output
"""
def format_transform_matrix(T):
r = R.from_matrix(T[:3, :3])
q = r.as_quat()
t = T[:3, 3]
return (t, q)
df_output['idx1'].append(idx1)
df_output['idx2'].append(idx2)
(t, q) = format_transform_matrix(T)
# translation:
df_output['t_x'].append(t[0])
df_output['t_y'].append(t[1])
df_output['t_z'].append(t[2])
# rotation:
df_output['q_w'].append(q[3])
df_output['q_x'].append(q[0])
df_output['q_y'].append(q[1])
df_output['q_z'].append(q[2])
def write_output(filename, df_output):
"""
Write output
"""
df_output = pd.DataFrame.from_dict(
df_output
)
print(f'write output to {filename}')
df_output[
[
'idx1', 'idx2',
't_x', 't_y', 't_z',
'q_w', 'q_x', 'q_y', 'q_z'
]
].to_csv(filename, index=False)
visualize.py
import numpy as np
import open3d as o3d
def show_inlier_outlier(cloud, ind):
"""
Visualize inliers and outliers
"""
inlier_cloud = cloud.select_by_index(ind)
outlier_cloud = cloud.select_by_index(ind, invert=True)
outlier_cloud.paint_uniform_color([1, 0, 0])
inlier_cloud.paint_uniform_color([0.5, 0.5, 0.5])
o3d.visualization.draw_geometries([inlier_cloud, outlier_cloud])
def get_point_cloud_diameter(pcd):
"""
Get point cloud diameter by min-max bounding box
"""
diameter = np.linalg.norm(
pcd.get_max_bound() - pcd.get_min_bound()
)
return diameter
def show_registration_result(
pcd_source_keypoints, pcd_target_keypoints, association,
pcd_source_dense, pcd_target_dense, transformation
):
"""
Visualize point cloud registration results.
Parameters
----------
pcd_source_keypoints: open3d.geometry.PointCloud
keypoints in source point cloud
pcd_target_keypoints: open3d.geometry.PointCloud
keypoints in target point cloud
association: numpy.ndarray
keypoint associations from feature matching
pcd_source_dense: open3d.geometry.PointCloud
filtered source point cloud
pcd_target_dense: open3d.geometry.PointCloud
filtered target point cloud
transformation: numpy.ndarray
transformation matrix
Returns
----------
None
"""
#
# group 01 -- registration result:
#
# apply transform:
pcd_source_dense.transform(transformation)
# move registration result to origin:
center_source, _ = pcd_source_dense.compute_mean_and_covariance()
center_target, _ = pcd_target_dense.compute_mean_and_covariance()
translation = 0.5 * (center_source + center_target)
pcd_source_dense_centered = pcd_source_dense.translate(-translation)
pcd_target_dense_centered = pcd_target_dense.translate(-translation)
# draw result:
pcd_source_dense_centered.paint_uniform_color([1, 0.706, 0])
pcd_target_dense_centered.paint_uniform_color([0, 0.651, 0.929])
#
# group 02 -- keypoint association result:
#
# get diameters of source and target:
diameter_source = get_point_cloud_diameter(pcd_source_dense)
diameter_target = get_point_cloud_diameter(pcd_target_dense)
# shift correspondence sets:
diameter = max(diameter_source, diameter_target)
pcd_source_keypoints_shifted = pcd_source_keypoints.translate(
-translation + np.asarray([diameter, -diameter, 0.0])
)
pcd_target_keypoints_shifted = pcd_target_keypoints.translate(
-translation + np.asarray([diameter, +diameter, 0.0])
)
# draw associations:
association = np.asarray(association)[:20,:]
N, _ = association.shape
# keep only 20 pairs:
points = np.vstack(
(
np.asarray(pcd_source_keypoints_shifted.points)[association[:, 0]],
np.asarray(pcd_target_keypoints_shifted.points)[association[:, 1]]
)
)
correspondences = np.asarray(
[
[i, i + N] for i in np.arange(N)
]
)
colors = [
[0.0, 0.0, 0.0] for i in range(N)
]
correspondence_set = o3d.geometry.LineSet(
points = o3d.utility.Vector3dVector(points),
lines = o3d.utility.Vector2iVector(correspondences),
)
correspondence_set.colors = o3d.utility.Vector3dVector(colors)
pcd_source_keypoints_shifted.paint_uniform_color([0.0, 1.0, 0.0])
np.asarray(pcd_source_keypoints_shifted.colors)[association[:, 0], :] = [1.0, 0.0, 0.0]
pcd_target_keypoints_shifted.paint_uniform_color([0.0, 0.0, 1.0])
np.asarray(pcd_target_keypoints_shifted.colors)[association[:, 1], :] = [1.0, 0.0, 0.0]
o3d.visualization.draw_geometries(
[
# registration result:
pcd_source_dense_centered, pcd_target_dense_centered,
# feature point association:
pcd_source_keypoints_shifted, pcd_target_keypoints_shifted,correspondence_set
]
)
iss.py
import heapq
import numpy as np
import pandas as pd
import open3d as o3d
def detect(point_cloud, search_tree, radius):
"""
Detect point cloud key points using Intrinsic Shape Signature(ISS)
Parameters
----------
point_cloud: Open3D.geometry.PointCloud
input point cloud
search_tree: Open3D.geometry.KDTree
point cloud search tree
radius: float
radius for ISS computing
Returns
----------
point_cloud: numpy.ndarray
Velodyne measurements as N-by-3 numpy ndarray
"""
# points handler:
points = np.asarray(point_cloud.points)
# keypoints container:
keypoints = {
'id': [],
'x': [],
'y': [],
'z': [],
'lambda_0': [],
'lambda_1': [],
'lambda_2': []
}
# cache for number of radius nearest neighbors:
num_rnn_cache = {
}
# heapq for non-maximum suppression:
pq = []
for idx_center, center in enumerate(
points
):
# find radius nearest neighbors:
[k, idx_neighbors, _] = search_tree.search_radius_vector_3d(center, radius)
# use the heuristics from NDT to filter outliers:
if k < 6:
continue
# for each point get its nearest neighbors count:
w = []
deviation = []
for idx_neighbor in np.asarray(idx_neighbors[1:]):
# check cache:
if not idx_neighbor in num_rnn_cache:
[k_, _, _] = search_tree.search_radius_vector_3d(
points[idx_neighbor],
radius
)
num_rnn_cache[idx_neighbor] = k_
# update:
w.append(num_rnn_cache[idx_neighbor])
deviation.append(points[idx_neighbor] - center)
# calculate covariance matrix:
w = np.asarray(w)
deviation = np.asarray(deviation)
cov = (1.0 / w.sum()) * np.dot(
deviation.T,
np.dot(np.diag(w), deviation)
)
# get eigenvalues:
w, _ = np.linalg.eig(cov)
w = w[w.argsort()[::-1]]
# add to pq:
heapq.heappush(pq, (-w[2], idx_center))
# add to dataframe:
keypoints['id'].append(idx_center)
keypoints['x'].append(center[0])
keypoints['y'].append(center[1])
keypoints['z'].append(center[2])
keypoints['lambda_0'].append(w[0])
keypoints['lambda_1'].append(w[1])
keypoints['lambda_2'].append(w[2])
# non-maximum suppression:
suppressed = set()
while pq:
_, idx_center = heapq.heappop(pq)
if not idx_center in suppressed:
# suppress its neighbors:
[_, idx_neighbors, _] = search_tree.search_radius_vector_3d(
points[idx_center],
radius
)
for idx_neighbor in np.asarray(idx_neighbors[1:]):
suppressed.add(idx_neighbor)
else:
continue
# format:
keypoints = pd.DataFrame.from_dict(
keypoints
)
# first apply non-maximum suppression:
keypoints = keypoints.loc[
keypoints['id'].apply(lambda id: not id in suppressed),
keypoints.columns
]
# then apply decreasing ratio test:
keypoints = keypoints.loc[
(keypoints['lambda_0'] > keypoints['lambda_1']) &
(keypoints['lambda_1'] > keypoints['lambda_2']),
keypoints.columns
]
# finally, return the keypoint in decreasing lambda 2 order:
keypoints = keypoints.sort_values('lambda_2', axis=0, ascending=False, ignore_index=True)
return keypoints