聚类算法概述(k-Means++/FCM/凝聚层次聚类/DBSCAN)
欢迎光临我的博客:HaoyuHu’s Blog
参考自初识聚类算法:K均值、凝聚层次聚类和DBSCAN,模糊聚类FCM算法。
近期做完了labmu的tunet3.0,总算有时间学习一些东西了。目前想学的有聚类分析、图像识别算法和计算机网络方面的知识。在暑假实习期间,开始着手游戏编程。
聚类的目的
将数据划分为若干个簇,簇内相似性大,簇间相似性小,聚类效果好。用于从数据中提取信息和规律。
聚类的概念
- 层次与划分:当允许存在子簇时,将数据按照层次划分,最终得到的是一颗树。树中包含的层次关系即为聚类划分的层次关系。各个子簇不重叠,每个元素都隶属于某个level的子簇中。
- 互斥、重叠与模糊:这个概念的核心在于,所有集合元素都不完全隶属于任何一个簇,而是按照一定隶属度归属于所有簇。对于任意一个元素,其隶属度和一般为1。
- 完全与部分:完全聚类要求所有数据元素都必须有隶属,而部分聚类则允许噪音存在,不隶属于任何簇。
簇的分类
- 明显分离:不同簇间任意元素距离都大于簇内元素距离。从图像上观察是明显分离类型的簇。
- 基于原型:任意元素与它所隶属的簇的簇中心(簇内元素集合的质心)的距离大于到其他簇中心的距离。
- 基于图:图中节点为对象,弧权值为距离。类似于明显分离的定义或基于原型的定义,只是用弧权值代替了人为规定的距离。
- 基于密度:基于密度的簇分类是较为常用,也是应用范围最为广泛的一种分类方法。元素的稠密程度决定了簇的分布。当存在并希望分辨噪声时,或簇形状不规则时,往往采用基于密度的簇分类。
常用的聚类分析算法
- 基本k均值:即k-means算法。簇的分类是基于原型的。用于已知簇个数的情况,且要求簇的形状基本满足圆形,不能区分噪声。
- 凝聚层次聚类:起初各个点为一个簇,而后按照距离最近凝聚,知道凝聚得到的簇个数满足用户要求。
- DBscan:基于密度和划分的聚类方法。
聚类算法的基本思想
(1) 基本k均值聚类(hard c-means, HCM)
方法很简单,首先给出初始的几个簇中心。将所有元素按照到簇中心最近的归属原则,归属到各个簇。然后对各个簇求解新的簇中心(元素集合质心)。重复上述步骤直到质心不再明显变化后,即完成聚类。
采用何种距离可按照数据性质或项目要求。距离的分类可以参考A-star算法概述及其在游戏开发中的应用分析中提到的曼哈顿距离、对角线距离、欧几里得距离等。实际上相当于求解一个全局状态函数的最小值问题,状态函数是各个元素到最近簇中心的距离之和。
该算法的特点有如下几点:
- 其一,不一定得到全局最优解,当初始簇中心不满足要求时,可能只能得到局部最优解,当然有学者通过一定的预处理使得得到的初始簇中心满足一定条件,从而能够得到全局最优解,并将方法名改为k-means++。
- 其二,不能排除噪声点对聚类的影响。
- 其三,要求簇形状接近圆形。
- 要求完全聚类的情况。
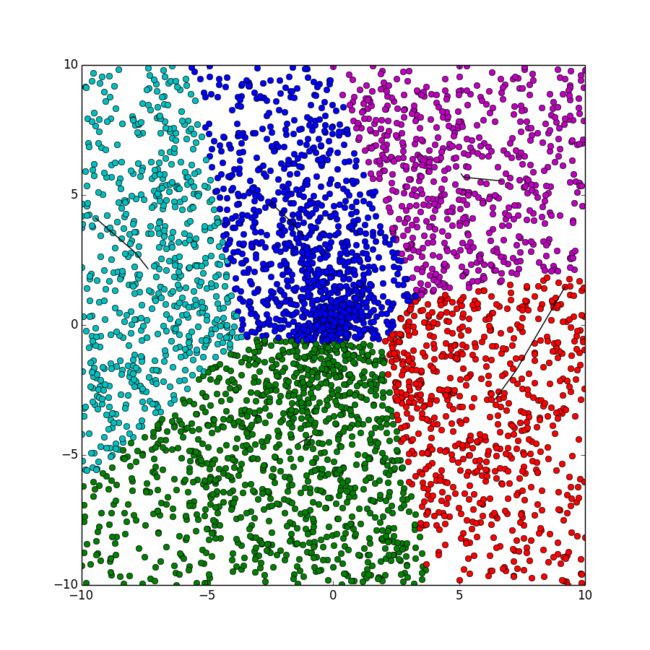
python代码
此代码使用的是k-means++算法,采用约定的方法使得到的初始聚类中心能够在后面的迭代过程中收敛到最优解。
import math
import collections
import random
import copy
import pylab
try:
import psyco
psyco.full()
except ImportError:
pass
FLOAT_MAX = 1e100
class Point:
__slots__ = ["x", "y", "group"]
def __init__(self, x = 0, y = 0, group = 0):
self.x, self.y, self.group = x, y, group
def generatePoints(pointsNumber, radius):
points = [Point() for _ in xrange(pointsNumber)]
for point in points:
r = random.random() * radius
angle = random.random() * 2 * math.pi
point.x = r * math.cos(angle)
point.y = r * math.sin(angle)
return points
def solveDistanceBetweenPoints(pointA, pointB):
return (pointA.x - pointB.x) * (pointA.x - pointB.x) + (pointA.y - pointB.y) * (pointA.y - pointB.y)
def getNearestCenter(point, clusterCenterGroup):
minIndex = point.group
minDistance = FLOAT_MAX
for index, center in enumerate(clusterCenterGroup):
distance = solveDistanceBetweenPoints(point, center)
if (distance < minDistance):
minDistance = distance
minIndex = index
return (minIndex, minDistance)
def kMeansPlusPlus(points, clusterCenterGroup):
clusterCenterGroup[0] = copy.copy(random.choice(points))
distanceGroup = [0.0 for _ in xrange(len(points))]
sum = 0.0
for index in xrange(1, len(clusterCenterGroup)):
for i, point in enumerate(points):
distanceGroup[i] = getNearestCenter(point, clusterCenterGroup[:index])[1]
sum += distanceGroup[i]
sum *= random.random()
for i, distance in enumerate(distanceGroup):
sum -= distance;
if sum < 0:
clusterCenterGroup[index] = copy.copy(points[i])
break
for point in points:
point.group = getNearestCenter(point, clusterCenterGroup)[0]
return
def kMeans(points, clusterCenterNumber):
clusterCenterGroup = [Point() for _ in xrange(clusterCenterNumber)]
kMeansPlusPlus(points, clusterCenterGroup)
clusterCenterTrace = [[clusterCenter] for clusterCenter in clusterCenterGroup]
tolerableError, currentError = 5.0, FLOAT_MAX
count = 0
while currentError >= tolerableError:
count += 1
countCenterNumber = [0 for _ in xrange(clusterCenterNumber)]
currentCenterGroup = [Point() for _ in xrange(clusterCenterNumber)]
for point in points:
currentCenterGroup[point.group].x += point.x
currentCenterGroup[point.group].y += point.y
countCenterNumber[point.group] += 1
for index, center in enumerate(currentCenterGroup):
center.x /= countCenterNumber[index]
center.y /= countCenterNumber[index]
currentError = 0.0
for index, singleTrace in enumerate(clusterCenterTrace):
singleTrace.append(currentCenterGroup[index])
currentError += solveDistanceBetweenPoints(singleTrace[-1], singleTrace[-2])
clusterCenterGroup[index] = copy.copy(currentCenterGroup[index])
for point in points:
point.group = getNearestCenter(point, clusterCenterGroup)[0]
return clusterCenterGroup, clusterCenterTrace
def showClusterAnalysisResults(points, clusterCenterTrace):
colorStore = ['or', 'og', 'ob', 'oc', 'om', 'oy', 'ok']
pylab.figure(figsize=(9, 9), dpi = 80)
for point in points:
color = ''
if point.group >= len(colorStore):
color = colorStore[-1]
else:
color = colorStore[point.group]
pylab.plot(point.x, point.y, color)
for singleTrace in clusterCenterTrace:
pylab.plot([center.x for center in singleTrace], [center.y for center in singleTrace], 'k')
pylab.show()
def main():
clusterCenterNumber = 5
pointsNumber = 2000
radius = 10
points = generatePoints(pointsNumber, radius)
_, clusterCenterTrace = kMeans(points, clusterCenterNumber)
showClusterAnalysisResults(points, clusterCenterTrace)
main()
(1)Extra 基于模糊数学的c均值聚类(FCM)
模糊c均值聚类(fuzzy c-means clustering)与硬划分k均值聚类相同,都是一种基于划分的聚类分析方法,但FCM是HCM的自然进阶版。与k均值聚类不同的是,模糊c均值聚类的点按照不同的隶属度ui隶属于不同的聚类中心vi,聚类的过程类似k均值聚类。(详见:模糊聚类FCM算法)
聚类步骤:
- 初始化。采用k-means++的方法确定初始聚类中心,确保最优解。
- 确定各个点对各个聚类中心的隶属度u(i,j)。m为加权指数。公式如下:
u(i,j) = (sum(distance(point(j), center(i)) / distance(point(j), center(k)))^(1/(m-1)))^-1- 确定新的聚类中心,标记聚类中心变化轨迹。公式如下:
v(i) = sum(u(i,j)^m * point(j)) / sum(u(i,j)^m)- 判断聚类中心变化幅值是否小于给定的误差限。如不满足返回步骤2,否则退出循环。
- 打印聚类中心轨迹和聚类结果。
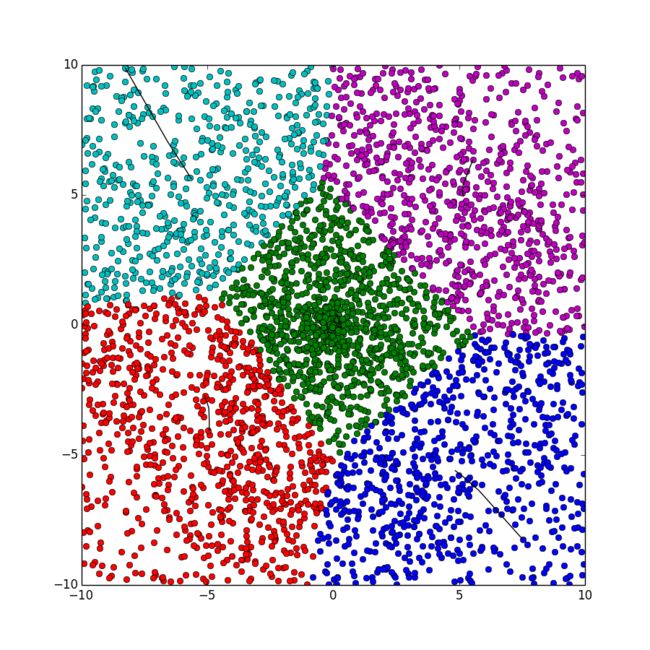
python代码
import math
import collections
import random
import copy
import pylab
try:
import psyco
psyco.full()
except ImportError:
pass
FLOAT_MAX = 1e100
class Point:
__slots__ = ["x", "y", "group", "membership"]
def __init__(self, clusterCenterNumber, x = 0, y = 0, group = 0):
self.x, self.y, self.group = x, y, group
self.membership = [0.0 for _ in xrange(clusterCenterNumber)]
def generatePoints(pointsNumber, radius, clusterCenterNumber):
points = [Point(clusterCenterNumber) for _ in xrange(2 * pointsNumber)]
count = 0
for point in points:
count += 1
r = random.random() * radius
angle = random.random() * 2 * math.pi
point.x = r * math.cos(angle)
point.y = r * math.sin(angle)
if count == pointsNumber - 1:
break
for index in xrange(pointsNumber, 2 * pointsNumber):
points[index].x = 2 * radius * random.random() - radius
points[index].y = 2 * radius * random.random() - radius
return points
def solveDistanceBetweenPoints(pointA, pointB):
return (pointA.x - pointB.x) * (pointA.x - pointB.x) + (pointA.y - pointB.y) * (pointA.y - pointB.y)
def getNearestCenter(point, clusterCenterGroup):
minIndex = point.group
minDistance = FLOAT_MAX
for index, center in enumerate(clusterCenterGroup):
distance = solveDistanceBetweenPoints(point, center)
if (distance < minDistance):
minDistance = distance
minIndex = index
return (minIndex, minDistance)
def kMeansPlusPlus(points, clusterCenterGroup):
clusterCenterGroup[0] = copy.copy(random.choice(points))
distanceGroup = [0.0 for _ in xrange(len(points))]
sum = 0.0
for index in xrange(1, len(clusterCenterGroup)):
for i, point in enumerate(points):
distanceGroup[i] = getNearestCenter(point, clusterCenterGroup[:index])[1]
sum += distanceGroup[i]
sum *= random.random()
for i, distance in enumerate(distanceGroup):
sum -= distance;
if sum < 0:
clusterCenterGroup[index] = copy.copy(points[i])
break
return
def fuzzyCMeansClustering(points, clusterCenterNumber, weight):
clusterCenterGroup = [Point(clusterCenterNumber) for _ in xrange(clusterCenterNumber)]
kMeansPlusPlus(points, clusterCenterGroup)
clusterCenterTrace = [[clusterCenter] for clusterCenter in clusterCenterGroup]
tolerableError, currentError = 1.0, FLOAT_MAX
while currentError >= tolerableError:
for point in points:
getSingleMembership(point, clusterCenterGroup, weight)
currentCenterGroup = [Point(clusterCenterNumber) for _ in xrange(clusterCenterNumber)]
for centerIndex, center in enumerate(currentCenterGroup):
upperSumX, upperSumY, lowerSum = 0.0, 0.0, 0.0
for point in points:
membershipWeight = pow(point.membership[centerIndex], weight)
upperSumX += point.x * membershipWeight
upperSumY += point.y * membershipWeight
lowerSum += membershipWeight
center.x = upperSumX / lowerSum
center.y = upperSumY / lowerSum
# update cluster center trace
currentError = 0.0
for index, singleTrace in enumerate(clusterCenterTrace):
singleTrace.append(currentCenterGroup[index])
currentError += solveDistanceBetweenPoints(singleTrace[-1], singleTrace[-2])
clusterCenterGroup[index] = copy.copy(currentCenterGroup[index])
for point in points:
maxIndex, maxMembership = 0, 0.0
for index, singleMembership in enumerate(point.membership):
if singleMembership > maxMembership:
maxMembership = singleMembership
maxIndex = index
point.group = maxIndex
return clusterCenterGroup, clusterCenterTrace
def getSingleMembership(point, clusterCenterGroup, weight):
distanceFromPoint2ClusterCenterGroup = [solveDistanceBetweenPoints(point, clusterCenterGroup[index]) for index in xrange(len(clusterCenterGroup))]
for centerIndex, singleMembership in enumerate(point.membership):
sum = 0.0
isCoincide = [False, 0]
for index, distance in enumerate(distanceFromPoint2ClusterCenterGroup):
if distance == 0:
isCoincide[0] = True
isCoincide[1] = index
break
sum += pow(float(distanceFromPoint2ClusterCenterGroup[centerIndex] / distance), 1.0 / (weight - 1.0))
if isCoincide[0]:
if isCoincide[1] == centerIndex:
point.membership[centerIndex] = 1.0
else:
point.membership[centerIndex] = 0.0
else:
point.membership[centerIndex] = 1.0 / sum
def showClusterAnalysisResults(points, clusterCenterTrace):
colorStore = ['or', 'og', 'ob', 'oc', 'om', 'oy', 'ok']
pylab.figure(figsize=(9, 9), dpi = 80)
for point in points:
color = ''
if point.group >= len(colorStore):
color = colorStore[-1]
else:
color = colorStore[point.group]
pylab.plot(point.x, point.y, color)
for singleTrace in clusterCenterTrace:
pylab.plot([center.x for center in singleTrace], [center.y for center in singleTrace], 'k')
pylab.show()
def main():
clusterCenterNumber = 5
pointsNumber = 2000
radius = 10
weight = 2
points = generatePoints(pointsNumber, radius, clusterCenterNumber)
_, clusterCenterTrace = fuzzyCMeansClustering(points, clusterCenterNumber, weight)
showClusterAnalysisResults(points, clusterCenterTrace)
main()
该算法的特点有如下几点:
- 主要特点与普通的k均值聚类类似。
- 要求完全聚类,不能区分噪声点。
- 聚类的中心符合度更高,但计算效率相对较低。
- 采用了平滑参数和隶属度的概念,使得各点的并不直接隶属于单个聚类中心。
(2) 凝聚层次聚类
初始状态各个元素各自为簇,每次合并簇间距离最小的簇。直到簇个数满足要求或合并超过90%。类似huffman树算法和查并集。上述距离的定义也有几种分类:包括簇间元素的最小距离,簇间元素的最大距离,和簇质心距离。
该算法的特点有如下几点:
- 凝聚聚类耗费的存储空间相对于其他几种方法要高。
- 可排除噪声点的干扰,但有可能噪声点分为一簇。
- 适合形状不规则,不要求聚类完全的情况。
- 合并操作不能撤销。
- 应注意,合并操作必须有一个合并限制比例,否则可能发生过度合并导致所有分类中心聚集,造成聚类失败。
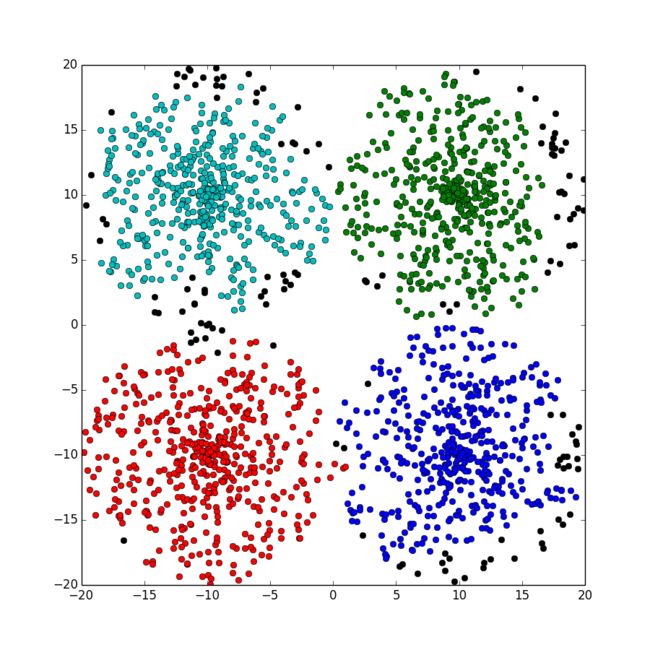
python代码
import math
import collections
import random
import copy
import pylab
try:
import psyco
psyco.full()
except ImportError:
pass
FLOAT_MAX = 1e100
class Point:
__slots__ = ["x", "y", "group"]
def __init__(self, x = 0, y = 0, group = 0):
self.x, self.y, self.group = x, y, group
def generatePoints(pointsNumber, radius):
points = [Point() for _ in xrange(4 * pointsNumber)]
originX = [-radius, -radius, radius, radius]
originY = [-radius, radius, -radius, radius]
count = 0
countCenter = 0
for index, point in enumerate(points):
count += 1
r = random.random() * radius
angle = random.random() * 2 * math.pi
point.x = r * math.cos(angle) + originX[countCenter]
point.y = r * math.sin(angle) + originY[countCenter]
point.group = index
if count >= pointsNumber * (countCenter + 1):
countCenter += 1
return points
def solveDistanceBetweenPoints(pointA, pointB):
return (pointA.x - pointB.x) * (pointA.x - pointB.x) + (pointA.y - pointB.y) * (pointA.y - pointB.y)
def getDistanceMap(points):
distanceMap = {}
for i in xrange(len(points)):
for j in xrange(i + 1, len(points)):
distanceMap[str(i) + '#' + str(j)] = solveDistanceBetweenPoints(points[i], points[j])
distanceMap = sorted(distanceMap.iteritems(), key=lambda dist:dist[1], reverse=False)
return distanceMap
def agglomerativeHierarchicalClustering(points, distanceMap, mergeRatio, clusterCenterNumber):
unsortedGroup = {index: 1 for index in xrange(len(points))}
for key, _ in distanceMap:
lowIndex, highIndex = int(key.split('#')[0]), int(key.split('#')[1])
if points[lowIndex].group != points[highIndex].group:
lowGroupIndex = points[lowIndex].group
highGroupIndex = points[highIndex].group
unsortedGroup[lowGroupIndex] += unsortedGroup[highGroupIndex]
del unsortedGroup[highGroupIndex]
for point in points:
if point.group == highGroupIndex:
point.group = lowGroupIndex
if len(unsortedGroup) <= int(len(points) * mergeRatio):
break
sortedGroup = sorted(unsortedGroup.iteritems(), key=lambda group: group[1], reverse=True)
topClusterCenterCount = 0
print sortedGroup, len(sortedGroup)
for key, _ in sortedGroup:
topClusterCenterCount += 1
for point in points:
if point.group == key:
point.group = -1 * topClusterCenterCount
if topClusterCenterCount >= clusterCenterNumber:
break
return points
def showClusterAnalysisResults(points):
colorStore = ['or', 'og', 'ob', 'oc', 'om', 'oy', 'ok']
pylab.figure(figsize=(9, 9), dpi = 80)
for point in points:
color = ''
if point.group < 0:
color = colorStore[-1 * point.group - 1]
else:
color = colorStore[-1]
pylab.plot(point.x, point.y, color)
pylab.show()
def main():
clusterCenterNumber = 4
pointsNumber = 500
radius = 10
mergeRatio = 0.025
points = generatePoints(pointsNumber, radius)
distanceMap = getDistanceMap(points)
points = agglomerativeHierarchicalClustering(points, distanceMap, mergeRatio, clusterCenterNumber)
showClusterAnalysisResults(points)
main()
(3) DBscan
DBscan是一种基于密度的聚类算法。因此首先应定义密度的概念。密度是以一个点为中心2*EPs边长的正方形区域内点的个数。并将不同密度的点划归为不同类型的点:
- 当密度大于阈值MinPs时,称为核心点。
- 当密度小于阈值MinPs,但领域内核心点的数量大于等于1,称为边界点。
- 非核心点且非边界点,称为噪声点。
具体操作:
- 将所有邻近的核心点划分到同一个簇中。
- 将所有边界点划分到其领域内的核心点的簇中。
- 噪声点不做归属处理。
该算法的特点有如下几点:
- 可排除噪声点的干扰。
- 适合形状不规则,不要求聚类完全的情况。
- 合并操作不能撤销。
minPointsNumberWithinBoundary和Eps决定了聚类的粒度和范围,当Eps增大或minPointsNumberWithinBoundary减小时,都会使聚类的粒度更粗,形成范围更大的簇。对于特定的问题,需要调整Eps和minPointsNumberWithinBoundary以满足聚类的要求。- 基于密度的聚类一定程度上回避了距离的计算,可以提高效率。
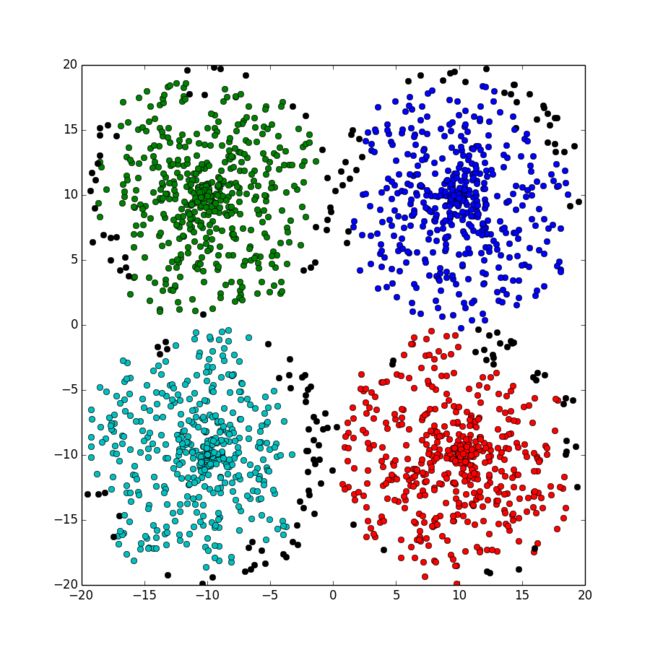
python代码
import math
import collections
import random
import copy
import pylab
try:
import psyco
psyco.full()
except ImportError:
pass
FLOAT_MAX = 1e100
CORE_POINT_TYPE = -2
BOUNDARY_POINT_TYPE = 1 #ALL NONE-NEGATIVE INTEGERS CAN BE BOUNDARY POINT TYPE
OTHER_POINT_TYPE = -1
class Point:
__slots__ = ["x", "y", "group", "pointType"]
def __init__(self, x = 0, y = 0, group = 0, pointType = -1):
self.x, self.y, self.group, self.pointType = x, y, group, pointType
def generatePoints(pointsNumber, radius):
points = [Point() for _ in xrange(4 * pointsNumber)]
originX = [-radius, -radius, radius, radius]
originY = [-radius, radius, -radius, radius]
count = 0
countCenter = 0
for index, point in enumerate(points):
count += 1
r = random.random() * radius
angle = random.random() * 2 * math.pi
point.x = r * math.cos(angle) + originX[countCenter]
point.y = r * math.sin(angle) + originY[countCenter]
point.group = index
if count >= pointsNumber * (countCenter + 1):
countCenter += 1
return points
def solveDistanceBetweenPoints(pointA, pointB):
return (pointA.x - pointB.x) * (pointA.x - pointB.x) + (pointA.y - pointB.y) * (pointA.y - pointB.y)
def isInPointBoundary(centerPoint, customPoint, halfScale):
return customPoint.x <= centerPoint.x + halfScale and customPoint.x >= centerPoint.x - halfScale and customPoint.y <= centerPoint.y + halfScale and customPoint.y >= centerPoint.y - halfScale
def getPointsNumberWithinBoundary(points, halfScale):
pointsIndexGroupWithinBoundary = [[] for _ in xrange(len(points))]
for centerIndex, centerPoint in enumerate(points):
for index, customPoint in enumerate(points):
if centerIndex != index and isInPointBoundary(centerPoint, customPoint, halfScale):
pointsIndexGroupWithinBoundary[centerIndex].append(index)
return pointsIndexGroupWithinBoundary
def decidePointsType(points, pointsIndexGroupWithinBoundary, minPointsNumber):
for index, customPointsGroup in enumerate(pointsIndexGroupWithinBoundary):
if len(customPointsGroup) >= minPointsNumber:
points[index].pointType = CORE_POINT_TYPE
for index, customPointsGroup in enumerate(pointsIndexGroupWithinBoundary):
if len(customPointsGroup) < minPointsNumber:
for customPointIndex in customPointsGroup:
if points[customPointIndex].pointType == CORE_POINT_TYPE:
points[index].pointType = customPointIndex
def mergeGroup(points, fromIndex, toIndex):
for point in points:
if point.group == fromIndex:
point.group = toIndex
def dbscan(points, pointsIndexGroupWithinBoundary, clusterCenterNumber):
countGroupsNumber = {index: 1 for index in xrange(len(points))}
for index, point in enumerate(points):
if point.pointType == CORE_POINT_TYPE:
for customPointIndex in pointsIndexGroupWithinBoundary[index]:
if points[customPointIndex].pointType == CORE_POINT_TYPE and points[customPointIndex].group != point.group:
countGroupsNumber[point.group] += countGroupsNumber[points[customPointIndex].group]
del countGroupsNumber[points[customPointIndex].group]
mergeGroup(points, points[customPointIndex].group, point.group)
#point.pointType >= 0 means it is BOUNDARY_POINT_TYPE
elif point.pointType >= 0:
corePointGroupIndex = points[point.pointType].group
countGroupsNumber[corePointGroupIndex] += countGroupsNumber[point.group]
del countGroupsNumber[point.group]
point.group = corePointGroupIndex
countGroupsNumber = sorted(countGroupsNumber.iteritems(), key=lambda group: group[1], reverse=True)
count = 0
for key, _ in countGroupsNumber:
count += 1
for point in points:
if point.group == key:
point.group = -1 * count
if count >= clusterCenterNumber:
break
def showClusterAnalysisResults(points):
colorStore = ['or', 'og', 'ob', 'oc', 'om', 'oy', 'ok']
pylab.figure(figsize=(9, 9), dpi = 80)
for point in points:
color = ''
if point.group < 0:
color = colorStore[-1 * point.group - 1]
else:
color = colorStore[-1]
pylab.plot(point.x, point.y, color)
pylab.show()
def main():
clusterCenterNumber = 4
pointsNumber = 500
radius = 10
Eps = 2
minPointsNumber = 18
points = generatePoints(pointsNumber, radius)
pointsIndexGroupWithinBoundary = getPointsNumberWithinBoundary(points, Eps)
decidePointsType(points, pointsIndexGroupWithinBoundary, minPointsNumber)
dbscan(points, pointsIndexGroupWithinBoundary, clusterCenterNumber)
showClusterAnalysisResults(points)
main()
后记
在学习和分析过程中发现几点待解决的问题:
- 其一,上述聚类过程都需要人为指定聚类中心数目,然而聚类的过程如果需人为干预,这可能是一个比较麻烦的问题。解决办法可以是采用多个候选聚类中心数目
{i,i+1,...k},对于不同的聚类中心数目都会有对应的分析结果,再采用贝叶斯定理。另一方面,机器无法知道人所需要的聚类粒度和聚类数目,如果完全由机器确定,也是不合理的。 - 其二,k-means聚类必须是完全聚类,对距离的选择也可以依据问题而定。
- 其三,实际上凝聚层次聚类和基于密度的dbscan聚类都有一个合并的过程,对于这种合并最好的算法应该是查并集,其时间复杂度为
O(n * f(n)),对于目前常见的大整数n,f(n) < 4。但如果过于追求效率,那么就违背了python语言开发和分析数据的优势。 - 其四,凝聚层次聚类和基于密度的dbscan聚类都对合并的程度有一定要求。凝聚层次聚类通过
mergeRatio来确定合并的比例;而dbscan是通过Eps和minPointsNumber来确定聚类的粒度。