sklearn预测pima糖尿病
文章目录
-
- 数据集描述
-
- 准备工作
- 实验环境和工具
- 预测分析
-
- 探索性数据分析
-
- 数据描述
- 可视化分析
- 构建baseline
- 数据预处理
-
- 离群值处理
- 缺失值处理
- 特征工程
- 数据标准化
- 模型构建与调参优化
- 完整代码
数据集描述
本数据集内含十个属性列
Pergnancies: 怀孕次数
Glucose:血糖浓度
BloodPressure:舒张压(毫米汞柱)
SkinThickness:肱三头肌皮肤褶皱厚度(毫米)
Insulin:两个小时血清胰岛素(μU/毫升)
BMI:身体质量指数,体重除以身高的平方
Diabets Pedigree Function: 疾病血统指数
是否和遗传相关,Height:身高(厘米)
Age:年龄
Outcome:0表示不患病,1表示患病。
任务:建立机器学习模型以准确预测数据集中的患者是否患有糖尿病
准备工作
查阅资料得知各属性的数据值要求,方面后期对于数据的分析与处理过程。
属性列名称 数据值要求
Pergnancies(怀孕次数) 符合常理即可(可为0)
Glucose(血糖浓度) 正常值为:80~120
BloodPressure(舒张压(毫米汞柱)) 正常值为:60~80
SkinThickness(肱三头肌皮肤褶皱厚度(毫米)) 不为0
Insulin(两个小时血清胰岛素(/毫升)) 正常值为:35~145
BMI(身体质量指数:体重除以身高的平方) 正常值为:18.5~24.9
Diabets Pedigree Function:(疾病血统指数:是否和遗传相关) 无特殊值要求
Height(身高(厘米)) 不为0 符合常理即可
Age(年龄) 符合常理即可
Outcome(0表示不患病,1表示患病) 标签值
实验环境和工具
python3.5.6 + jupyter
数据处理 pandas、numpy
可视化 matplotlib、seaborn
模型构建 sklearn
预测分析
探索性数据分析
数据描述
首先观察基本的数据类型,以及数据是否存在缺失情况,简要统计信息
all_data.shape
all_data.info()
<class 'pandas.core.frame.DataFrame'>
RangeIndex: 768 entries, 0 to 767
Data columns (total 10 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 Pregnancies 768 non-null int64
1 Glucose 768 non-null int64
2 BloodPressure 768 non-null int64
3 SkinThickness 768 non-null int64
4 Insulin 768 non-null int64
5 BMI 768 non-null float64
6 DiabetesPedigreeFunction 768 non-null float64
7 Age 768 non-null int64
8 Height 766 non-null object
9 Outcome 768 non-null int64
dtypes: float64(2), int64(7), object(1)
memory usage: 60.1+ KB
数据总量时比较少的只有768个例子,可以看到除Height外的属性都为数值型属性。在后续数据预处理过程需要对Height属性进行类型转换操作。目前没有缺失值的出现。
# height 数值类型 为object 需要转化为 数值型
all_data = all_data.astype({
'Height':'float64'})
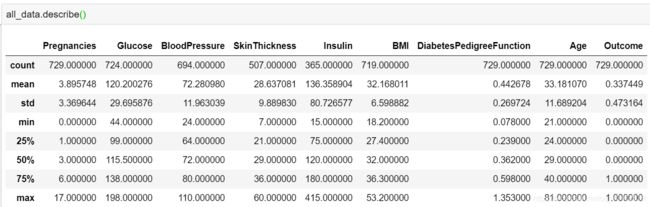
all_data.describe()
- 缺失值
从其中的min值可以很直观地观察到,Glucose, BloodPressure, SkinTinckness, Insulin, BMI等特征存在0值的情况(当然Pregnancies根据常识判断是可以为0的)。而根据常规范围明显可以判定这些0值是不合理的,所以也是一种缺失值缺失值,后续数据预处理需要对这些缺失值进行填充处理。 - 离群值/异常值
Glucose,BloodPressure,SkinTinckness,Insulin等特征的max值和75%分位点值或者min值和25%分位点值之间的差距比较大,初步判断可能存在离群值/异常值。尤其是SkinThickness和Insulin特征(具体见图4红色框部分),后续可以通过可视化进一步直观地观察判断。
为了方便后序对缺失值的统一处理,将异常值统一替换为np.nan。
import numpy as np
#缺失值替换 经分析,除怀孕次数,其他特征的0值表示缺失值 替换为np.nan
replace_list = ['Glucose', 'BloodPressure', 'SkinThickness', 'Insulin', 'BMI', 'Height']
all_data.loc[:,replace_list] = all_data.loc[:,replace_list].replace({
0:np.nan})
#各特征缺失数量统计
null_count = all_data.isnull().sum().values
# 缺失值情况
plt.figure()
sns.barplot(x = null_count, y = all_data.columns)
for x, y in enumerate(null_count):
plt.text(y, x, "%s" %y, horizontalalignment='center', verticalalignment='center')
plt.show()
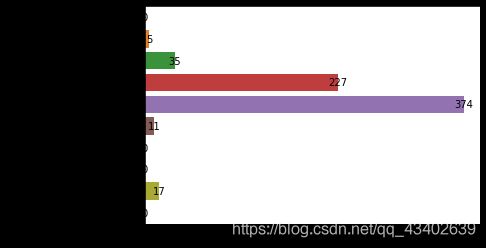
可以观察到Glucose,Insulin,SkinThickness,BMI,Height等特征都存在缺失值。并且 Insulin,SkinThickness缺失值比较多,分别占到了48%,30%的比例。所以后期数据预处理也是很关键的。
可视化分析
接下来通过更多针对性的可视化,来进一步探索特征值的分布以及特征和预测变量之间的关系
# 患病和不患病情况下 箱线图查看数据分散情况
for col in all_data.columns:
plt.figure(figsize = (10,6))
if all_data[col].unique().shape[0] > 2:
sns.boxplot(x="Outcome", y=col, data=all_data.dropna())
else:
sns.countplot(col,hue = 'Outcome',data = all_data.dropna())
plt.title(col)
plt.show()
观察患病和不患病情况下 各特征值或者人数分布
label接近2:1 存在一定的分布不平衡
像insulin之类的特征离群值是比较多的,由于离群值会对模型评估产生影响,所以后续可能要做处理,剔除偏离较大的离群值
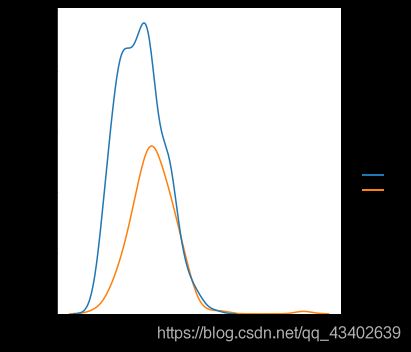
# 患病和不患病情况下 各特征的分布情况
for col in all_data.drop('Outcome',1).columns:
plt.figure()
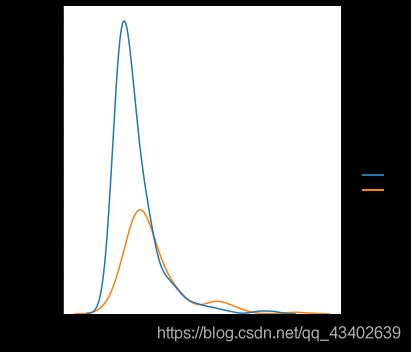
sns.displot(data = all_data, x = col,hue = 'Outcome',kind='kde')
plt.show()
- 从数据样本本身出发研究数据分布特征,可以发现在患病和不患病两种情况下,部分特征的密度分布比较相近,特别是height的分布图,发现两曲线基本相近。感觉和label之间的相关性都不是很强。
- 同时,可以发现部分特征存在右偏的现象(skewness (偏度) 描述数据分布形态的统计量,其描述的是某总体取值分布的对称性),考虑到需要数据尽量服从正态分布,所以后续数据预处理需要对存在一定偏度的特征进行相关处理。
# 观察各特征分布和患病的关系
corr = all_data.corr()
plt.figure(figsize = (8,6))
sns.heatmap(corr,annot = True,cmap = 'Blues')
plt.show()

heatmap()函数可以直观地将数据值的大小以定义的颜色深浅表示出来。
- 可以发现颜色相对来说都比较浅,也就是说无论是特征和特征之间还是特征和outcome标签之间的相关性都没有很高。
- 发现其余各特征变量中与outcome的相关度中最高的是Glucose 属性值为0.49,最低的是Height属性值为0.059。
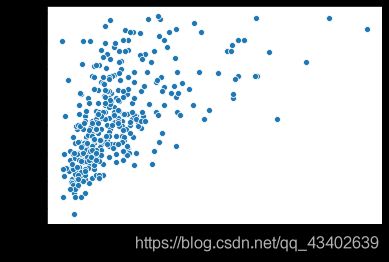
- 同时观察特征与特征之间的关系,发现Insulin与Glucose,BMI和SkinThickness之间的相关度分别为0.58,0.65属于比较高的相关性,由于Insulin是一个确实比较严重的特征,而相关性可以是一种协助填充缺失值的方法。
plt.figure()
sns.scatterplot(x = 'Insulin', y = 'Glucose', data = all_data)
plt.show()
sns.scatterplot(x = 'Insulin', y = 'BMI', data = all_data)
plt.show()
sns.scatterplot(x = 'Insulin', y = 'Age', data = all_data)
plt.show()
plt.figure()
sns.scatterplot(x = 'SkinThickness', y = 'BMI', data = all_data)
plt.show()
sns.scatterplot(x = 'SkinThickness', y = 'Glucose', data = all_data)
plt.show()
sns.scatterplot(x = 'SkinThickness', y = 'BloodPressure', data = all_data)
plt.show()
构建baseline
因为决策树几乎不需要数据预处理。其他方法经常需要数据标准化,创建虚拟变量和删除缺失值。
# 读取数据
all_data = pd.read_csv('data.csv')
# height 数值类型 为object 需要转化为 数值型
all_data = all_data.astype({
'Height':'float64'})
#
all_data.dropna(inplace = True)
# 特征
feature_data = all_data.drop('Outcome',1)
# 标签
label = all_data['Outcome']
base_model = DecisionTreeClassifier()
base_scores = cross_validate(base_model, feature_data, label,cv=5,return_train_score=True)
print(base_scores['test_score'].mean())
0.6954248366013072
数据预处理
综合前面分析,先做了以下处理
# 读取数据
all_data = pd.read_csv('data.csv')
# height 数值类型 为object 需要转化为 数值型
all_data = all_data.astype({
'Height':'float64'})
# 理论缺失值0替换为np.nan
replace_list = ['Glucose', 'BloodPressure', 'SkinThickness', 'Insulin', 'BMI', 'Height']
all_data.loc[:,replace_list] = all_data.loc[:,replace_list].replace({
0:np.nan})
# 删除相关性低的Height
all_data.drop('Height',1,inplace = True)
离群值处理
- 经过前面的分析发现数据是存在部分离群值的,虽然实验本身就是关于疾病预测,异常值的存在属于正常现象。但是对于一些可能超出人体接受范围的值,衡量对预测的影响之后,由于数据量比较小,这里选择删除极端异常点。
- 极端异常点 :上限的计算公式为Q3+3(Q3-Q1) 下界的计算公式为Q1-3(Q3-Q1))。
# remove the outliers
# 异常点 上须的计算公式为Q3+1.5(Q3-Q1);下须的计算公式为Q1-1.5(Q3-Q1)
# 极端异常点 :上限的计算公式为Q3+3(Q3-Q1) 下界的计算公式为Q1-3(Q3-Q1)
# 由于数据量比较少 所以选择删除极端异常值
def remove_outliers(feature,all_data):
first_quartile = all_data[feature].describe()['25%']
third_quartile = all_data[feature].describe()['75%']
iqr = third_quartile - first_quartile
# 异常值下标
index = all_data[(all_data[feature] < (first_quartile - 3*iqr)) | (all_data[feature] > (first_quartile + 3*iqr))].index
all_data = all_data.drop(index)
return all_data
outlier_features = ['Insulin', 'Glucose', 'BloodPressure', 'SkinThickness', 'BMI', 'DiabetesPedigreeFunction']
for feat in outlier_features:
all_data = remove_outliers(feat,all_data)
缺失值处理
缺失值处理这里考虑
- 直接删除处理
def drop_method(all_data):
median_fill = ['Glucose', 'BloodPressure','SkinThickness', 'BMI','Height']
for column in median_fill:
median_val = all_data[column].median()
all_data[column].fillna(median_val, inplace=True)
all_data.dropna(inplace = True)
return all_data
- 中值填充
def median_method():
for column in list(all_data.columns[all_data.isnull().sum() > 0]):
median = all_data[column].median()
all_data[column].fillna(median, inplace=True)
- KNNImputer填充
def knn_method():
# 先将缺失值比较少的特征用中值填充
values = {
'Glucose': all_data['Glucose'].median(),'BloodPressure':all_data['BloodPressure'].median(),'BMI':all_data['BMI'].median()}
all_data.fillna(value=values,inplace=True)
# 用KNNImputer 填充 Insulin SkinThickness
corr_SkinThickness = ['BMI', 'Glucose','BloodPressure', 'SkinThickness']
# 权重按距离的倒数表示。在这种情况下,查询点的近邻比远处的近邻具有更大的影响力
SkinThickness_imputer = KNNImputer(n_neighbors = 16,weights = 'distance')
all_data[corr_SkinThickness] = SkinThickness_imputer.fit_transform(all_data[corr_SkinThickness])
corr_Insulin = ['Glucose', 'BMI','BloodPressure', 'Insulin']
Insulin_imputer = KNNImputer(n_neighbors = 16,weights = 'distance')
all_data[corr_Insulin] = Insulin_imputer.fit_transform(all_data[corr_Insulin])
- 随机森林填充
from sklearn.ensemble import RandomForestRegressor
from sklearn.impute import SimpleImputer # 用来填补缺失值
def predict_method(feature):
# 复制一份数据 避免对原数据做出不必要的修改
copy_data = all_data.copy()
# 缺失了的下标
predict_index = copy_data[copy_data[feature].isnull()].index
# 没缺失的下标
train_index = copy_data[feature].dropna().index
# 用作预测 的训练集标签
train_label = copy_data.loc[train_index,feature]
copy_data = copy_data.drop(feature,axis=1)
# 对特征先用中值填充
imp_median = SimpleImputer(strategy='median')
# 用作预测的训练集特征
train_feature = copy_data.loc[train_index]
train_feature = imp_median.fit_transform(train_feature)
# 需要进行预测填充处理的缺失值
pre_feature = copy_data.loc[predict_index]
pre_feature = imp_median.fit_transform(pre_feature)
# 选取随机森林模型
fill_model = RandomForestRegressor()
fill_model = fill_model.fit(train_feature,train_label)
# 预测 填充
pre_value = fill_model.predict(pre_feature)
all_data.loc[predict_index,feature] = pre_value
#用随机森林的方法填充缺失值较多的 SkinThickness 和 Insulin 缺失值
predict_method("Insulin")
predict_method("SkinThickness")
# 其余值中值填充
for column in list(all_data.columns[all_data.isnull().sum() > 0]):
median = all_data[column].median()
all_data[column].fillna(median, inplace=True)
特征工程
# 特征
feture_data = all_data.drop('Outcome',1)
# 标签
label = all_data['Outcome']
# 利用BMI和身高构造weight特征
# BMI = weight(kg) / height(m)**2
feture_data['weight'] = (0.01*feture_data['Height'])**2 * feture_data['BMI']
数据标准化
# 标准化
Std = StandardScaler()
feture_data = Std.fit_transform(feture_data)
模型构建与调参优化
用到的模型
from sklearn.svm import SVC,SVR
from sklearn.tree import DecisionTreeClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier,StackingClassifier
调参方法
from sklearn.model_selection import GridSearchCV
def train(model, params):
grid_search = GridSearchCV(estimator = model, param_grid = params, cv = kfold)
grid_search.fit(feture_data,label)
print(grid_search.best_params_)
model_score = cross_validate(grid_search.best_estimator_,feture_data, label, cv=5)
print(model_score['test_score'])
print("mean test score :{}".format(model_score['test_score'].mean()))
return grid_search
SVC
#调参时先尝试一个大范围,确定比较小的范围,然后在小范围里搜索
model = SVC()
params = {
'C':np.linspace(0.1, 2, 100)}
SVC_grid_search = train(model,params)
plt.figure()
sns.lineplot(x=[x for x in range(100)],y=SVC_grid_search.cv_results_['mean_test_score'])
plt.show()
LogisticRegression
params = {
"C":np.linspace(0.1,2,100)}
model = LogisticRegression()
LR_grid_search= train(model,params)
plt.figure()
sns.lineplot(x=[x for x in range(100)],y=LR_grid_search.cv_results_['mean_test_score'])
plt.show()
RandomForestClassifier
params = {
"n_estimators":[x for x in range(30,50,4)],'min_samples_split':[x for x in range(2,12)]}
model = RandomForestClassifier()
RFC_grid_search = train(model,params)
plt.figure()
sns.lineplot(x=[x for x in range(len(grid_search.cv_results_['mean_test_score']))],
y=RFC_grid_search.cv_results_['mean_test_score'])
plt.show()
StackingClassifier
estimators = [
('SVC',SVC_grid_search.best_estimator_),
('NB', LR_grid_search.best_estimator_),
('RFC', RFC_grid_search.best_estimator_)
]
model = StackingClassifier(estimators=estimators, final_estimator=SVC())
model_score = cross_validate(model,feture_data, label, cv=5)
print(model_score['test_score'])
print("mean test score :{}".format(model_score['test_score'].mean()))
缺失值直接删除预测结果:
{‘C’: 1.405050505050505}
[0.83333333 0.71830986 0.83098592 0.83098592 0.84507042]
mean test score :0.811737089201878
{‘C’: 0.17676767676767677}
[0.86111111 0.73239437 0.77464789 0.83098592 0.84507042]
mean test score :0.8088419405320814
{‘min_samples_split’: 7, ‘n_estimators’: 30}
[0.77777778 0.69014085 0.74647887 0.83098592 0.85915493]
mean test score :0.780907668231612
[0.84722222 0.73239437 0.81690141 0.84507042 0.85915493]
mean test score :0.8201486697965571
缺失值中值填充预测效果
{‘C’: 1.7888888888888888}
[0.79452055 0.75342466 0.78082192 0.82191781 0.79310345]
mean test score :0.7887576759565423
{‘C’: 0.1575757575757576}
[0.78082192 0.76712329 0.7739726 0.80821918 0.77931034]
mean test score :0.7818894662257911
{‘min_samples_split’: 4, ‘n_estimators’: 44}
[0.80136986 0.71232877 0.74657534 0.81506849 0.79310345]
mean test score :0.7736891828058574
其余略 可以看出由于缺失值比较多,所以填充比直接删除的效果是要更差的
完整代码
https://github.com/wang-hui-shan/Pima_Diabetes_Predict